| Research Article | ||
Open Vet. J.. 2025; 15(2): 731-737 Open Veterinary Journal, (2025), Vol. 15(2): 731-737 Research Article Molecular study of Streptococcus equi isolated from horses with strangles in IraqSaif Aldeen Kamal Abdul-Latif* and Afaf Abdulrahman YousifDepartment of Internal and Preventive Veterinary Medicine, College of Veterinary Medicine, University of Baghdad, Baghdad, Iraq * Correspondence to: Saif Aldeen Kamal Abdul-Latif. Department of Internal and Preventive Veterinary Medicine, College of Veterinary Medicine, University of Baghdad, Baghdad, Iraq. saires27 [at] gmail.com Submitted: 24/10/2024 Accepted: 27/01/2025 Published: 28/02/2025 © 2025 Open Veterinary Journal
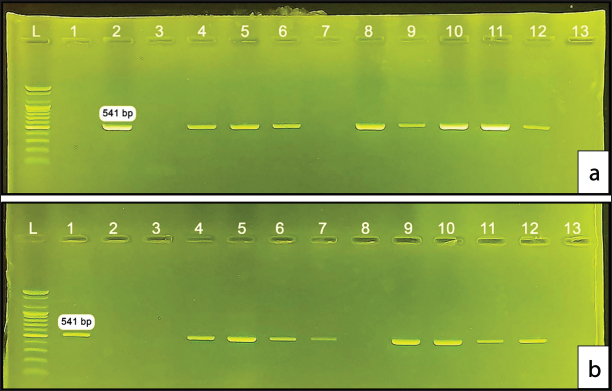
AbstractBackground: Strangles is a highly contagious equine respiratory disease caused by Streptococcus equi subsp. equi. It is a globally significant pathogen and one of the most common infectious agents in horses. In Iraq, no sequencing data on this pathogen are available, and only two molecular studies have been published to date. This study provides preliminary insights into strain diversity and provides a foundation for future large-scale investigations. Aim: his study aimed to investigate the molecular characteristics, identify SeM gene alleles, and perform a phylogenetic analysis of S. equi isolates from horses in Baghdad, Iraq. Methods: We analyzed 59 Streptococcus spp. isolates previously obtained from horses clinical samples. Conventional PCR (Polymerase Chain Reaction) targeting the SeM gene was used to identify S. equi. Additionally, nine PCR- positive SeM gene products were sequenced, followed by phylogenetic analysis and allele identification. Results: We confirmed 49 isolates as S. equi from the 59 isolates according to the molecular assay. Additionally, nine PCR products were used for sequencing and allele typing of the SeM gene, which provided the initial report of SeM-97 allele identification in Iraq. Phylogenetic analysis along with SeM gene typing revealed a close relationship between the Iraqi strains and one Iranian strain with 100% sequence identity, revealing important epidemiological relationships that may indicate regional ties to the strain detected in Iran. Conclusion: The present study represents the first investigation of SeM allele typing in Iraq, identifying the SeM-97 allele of S. equi along with its unique amino acid variations. The findings highlight genetic similarities between Iraqi isolates and a strain from Iran, suggesting the potential regional dissemination of S. equi. Keywords: Strangles, Streptococcus equi subsp. equi, PCR, SeM gene, SeM-97 IntroductionStrangles is a highly infectious and contagious bacterial infection of the upper respiratory system of horses and other equids of all ages and breeds, with high morbidity and low mortality in susceptible populations (Jaramillo-Morales et al., 2023). It is characterized by abrupt pyrexia followed by pharyngitis and subsequent abscess formation in the submandibular and retropharyngeal lymph nodes, and typically, young horses are frequently affected (Sweeney et al., 2005; Boyle, 2016; Arafa et al., 2021; Bekele et al., 2024). The causative organism, Streptococcus equi subsp. equi (S. equi), is a member of Lancefield Group C. It is a gram-positive, pus-forming bacteria that produces hemolysis on blood agar and is one of the most common infectious agents in horses (Sweeney et al., 1989, 2005; Holden et al., 2009; McGlennon et al., 2021). Strangles may be misdiagnosed and complicated by other infections, such as pneumonia, guttural pouch empyema, and purpura hemorrhagica (Whelchel and Chaffin, 2009; Arafa et al., 2024). The organism has several virulence factors shielding it from the host’s immune system, such as a hyaluronic acid capsule, superantigens, and M-like proteins SeM (Anzai et al., 1999; Artiushin et al., 2002; Proft et al., 2003; Timoney et al., 2014). SeM protein is responsible for increased virulence of S. equi over its ancestor, Streptococcus zooepidemicus, by actively binding to fibrinogen and IgG and inhibiting C3b deposition on the bacterial surface, triggering an antiphagocytic action resembling that of the M proteins of group A streptococci (Boschwitz and Timoney, 1994; Meehan et al., 2000; Boyle et al., 2009; Quinn et al., 2011). Serological tests, such as SeM-based and dual-target Enzyme-Linked Immunosorbent Assays (ELISAs), detect antibodies against S. equi and identify recent exposures and complications like purpura hemorrhagica (Boyle et al., 2009; Robinson et al., 2013). However, they cannot confirm active infection or carrier status, and cross-reactivity with S. zooepidemicus can limit their specificity (Duran and Goehring, 2021; Durham and Kemp-Symonds, 2021). The culture and biochemical identification of S. equi methods is of low sensitivity, whereas the molecular identification of S. equi based on the amplification and sequencing of the SeM gene has been used previously, with high sensitivity and specificity compared to culture (Newton et al., 1997; Timoney and Artiushin, 1997; Gronbaek et al., 2006). DNA sequencing of the hypervariable N-terminal region of the S. equi SeM gene is a useful tool for strain- typing to determine the source of strangles outbreaks (Kelly et al., 2006; Ivens et al., 2011; Kasuya et al., 2019; Rotinsulu et al., 2023). Streptococcus equi SeM allele typing has been well documented in China (Dong et al., 2019; Liu, 2019), Thailand (Tonpitak et al., 2016), New Zealand (Patty and Cursons, 2014), Ireland (Moloney et al., 2013), Brazil (Libardoni et al., 2013), Sweden (Lindahl et al., 2011), and Europe (Kelly et al., 2006). In Iraq, strangles remain a significant equine health concern, yet studies on their prevalence and molecular characterization are limited. A seroprevalence study in Mosul reported a 12% prevalence of anti-S. equi antibodies in horses, with higher rates observed in animals under 3 years old (20%) and those exhibiting respiratory signs (Al-Robaiee et al., 2023). In Baghdad, Mahmood (Mahmood et al., 2014) detected S. equi in 30 out of 141 clinical samples from horses with respiratory infections using conventional PCR, whereas Alwan (Alwan, 2014) employed Real-Time PCR targeting the SeM gene and identified S. equi in 24 out of 30 isolates (80%). Earlier, Jawad (Jawad, 2000) documented clinical cases of strangles in young horses and emphasized the potential of vaccination to control strangles. Similarly, Kaisium (Kaisium and Al-Judi, 2007) demonstrated the efficacy of a live attenuated vaccine in inducing immunity in rabbits. Despite these efforts, molecular studies on S. equi in Iraq remain scarce, with the last investigations conducted nearly a decade ago. Moreover, none of the previous studies examined the genetic diversity of S. equi or performed SeM gene sequencing, which is critical for strain typing and understanding the epidemiology of strangles. Strangles in Iraq likely originate from neighboring countries, such as Turkey and Iran, because of the uncontrolled and unsupervised importation of horses. The S. equi isolates from Iraq are hypothesized to have close genetic relationships with those from these regions. In addition, the SeM gene alleles circulating in Baghdad may exhibit significant variability, reflecting the diverse origins of the imported strains. This study aimed to molecularly identify S. equi isolates from horses with strangles in Baghdad, Iraq, sequence the SeM gene, identify circulating alleles, and perform phylogenetic comparisons with international strains. Materials and MethodsSource of Streptococcus isolatesFifty-nine isolates of Streptococcus spp. were obtained from the Department of Internal and Preventive Veterinary Medicine Laboratory, College of Veterinary Medicine, University of Baghdad. These isolates were collected during clinical practice on horses in Baghdad city and were initially identified as Streptococcus spp. using bacteriological methods. The identification included culturing on Columbia blood agar (Oxoid, UK) supplemented with 5% sheep blood for 24 hour at 37°C, Gram staining, and finally, suspected colonies purified by sub-culturing again on Columbia blood agar. Subsequently, biochemical identification was performed according to the sugar fermentation abilities of lactose, sorbitol, trehalose, and maltose in addition to the catalase test. These isolates were subsequently used in this study to identify S. equi via molecular methods. DNA extractionDNA was extracted and purified using Presto™ Minig DNA Bacteria Kit (Geneaid, Taiwan) according to the manufacturer’s instructions. The eluted DNA was stored at –20ºC until use. Polymerase chain reactionConventional PCR was used to amplify a 541 bp segment of the N-terminal of the SeM gene using the primers SeM-Forward ASW73 (5’-CAGAAAACTAAGTGCCGGTG-’3) and SeM-Reverse ASW74 (5’-ATTCGGTAAGAGCTTGACGC-’3) designed by (Kelly et al., 2006), in addition to GoTaq® Green Master Mix (Promega, USA), the thermocycler initial denaturation was set at 95°C for 5 minutes, denaturation at 90°C for 30 seconds, annealing at 55°C for 30 seconds, extension at 72°C for 30 minutes, and final extension at 72°C for 7 minutes. Negative control for PCRTo ensure the reliability of the results and monitor for potential contamination during DNA extraction and PCR amplification, a negative control was included in the procedure (Baverud et al., 2007; Bekele et al., 2024; Zu et al., 2024). The negative control was prepared during the DNA extraction process and consisted of only Gram+ buffer from the DNA extraction kit, with no sample added. This control underwent the entire DNA extraction process and PCR thermocycling alongside the test samples. By including this control, any contamination introduced during the extraction or amplification process could be identified and mitigated, ensuring the validity of the results obtained from the experimental samples. Sequencing of the SeM gene and data analysisPositive PCR amplifications with SeM-Forward primer were submitted for Sanger sequencing with ABI3730XL, an automated DNA sequencing platform, at Macrogen Corporation in Korea. The results were processed using Geneious Prime® and MEGA11 software. ResultsDNA extractionThe DNA of 59 suspected Streptococcus spp. isolates was successfully extracted, which appeared as compact bands on gel electrophoresis. The negative control showed no bands, confirming the absence of contamination during the extraction process. The concentration and purity of extracted DNA were measured by NanoDrop™ spectrophotometer, and they ranged between 88.8 and 141.5 ng/µl and 1.72 to 1.83 at absorbance 260/280 nm, respectively. PCR analysis of S. equiPCR targeting the SeM gene identified S. equi in 49 out of the 59 isolates (83.1%) based on the expected amplification product of 541 bp. This result confirms the presence of the causative agent of strangles in most of the tested samples (Fig. 1). Analysis of SeM gene sequencesNine random positive PCR products of the SeM gene were sequenced, and the data were utilized for further analysis. DNA sequence annotation and quality checks were conducted using Geneious Prime®. Next, nucleotide sequences were compared using the National Center for Biotechnology Information’s Basic Local Alignment Search Tool (BLAST) to the S. equi SeM gene, which was available in the GenBank database. BLAST results gave 188 significant alignments, and only one exact match was found (100%) for our SeM sequences query, and it was Iranian in origin, with (GenBank ID OL332314.1). Additionally, the nine Iraqi SeM gene sequences were submitted to the NCBI database, and accession numbers were assigned (PP726110.1, PP738564.1, PP738563.1, PP738562.1, PP738561.1, PP738560.1, PP738559.1, PP738558.1, PP738557.1). Eighteen NCBI SeM sequences from different countries were selected and downloaded; they had query coverage (100%) and an E-value of (0.0) compared to our sequences. Furthermore, the total number of 27 SeM sequences were aligned using the Multiple Sequence Comparison by Log-Expectation (MUSCLE) multiple alignment method (Edgar, 2004; Katoh and Standley, 2013) in Geneious Prime® 2024.0.5 software. A maximum likelihood tree was generated by PhyML version 3.3.2 (Guindon et al., 2010) using likelihood settings from the best-fit model (T93+G) selected by MEGA11 (Nei and Kumar, 2000; Tamura et al., 2021) and visualized using iTOL (Letunic and Bork, 2024), as shown in (Fig. 2). The result shows that the nine Iraqi strains, in addition to the Iranian strain (OL332314.1) (Moghaddam et al., 2023), had a close relationship with two other previously isolated S. equi strains from the UK (AJ249868.1) and Iran (MZ292707.1), with (99.79%) identity for both strains.
Fig. 1. (a) and (b) Electrophoreses of the agarose gel (1.2%) show the amplification of 541 bp SeM gene fragments from S. equi. Lane L shows the PCR ladder, and lane 13 shows the negative control.
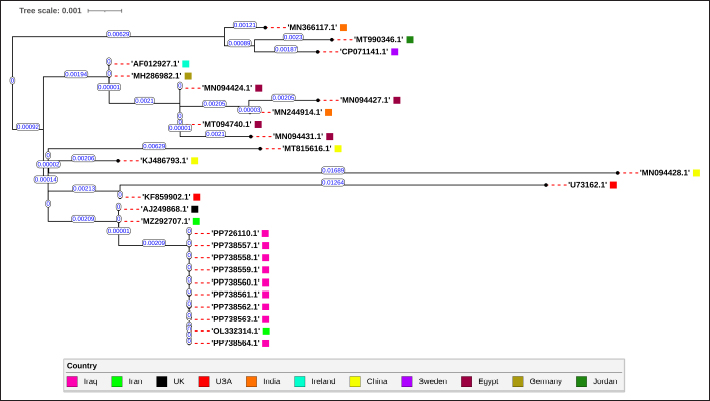
Fig. 2. Maximum likelihood tree illustrating the phylogenetic relationships among 27 different SeM gene sequences of S. equi, generated using PhyML version 3.3.2. The nine Iraqi strains (highlighted in pink), along with a highly similar Iranian strain, form a distinct cluster. This cluster shows a close evolutionary relationship with another group consisting of UK and Iranian strains. The scale bar represents the number of substitutions per site, indicating the degree of genetic divergence among sequences. Table 1. Comparison of the predicted amino acid sequences from the transcript of Iraqi S. equi and reference strain 4047.
SeM alleles identificationThe nine Iraqi SeM sequences were compared to the SeM allelic database in PubMLST. A single exact match was discovered, identifying the SeM-97 allele, which was previously isolated from a strain in Iran (OL332314.1) (Moghaddam et al., 2023). For evaluation of SeM gene alleles, a comparison of the predicted amino acid sequences of SeM-97 of the nine Iraqi strains to SeM-3 of the NCBI reference S. equi strain 4047 (NC_012471.1) (Holden et al., 2009) identified 4 amino acid differences (Table 1). DiscussionThis study provides foundational insights into the genetic variability and strain relationships of S. equi in Iraq while acknowledging the need for larger scale investigations to address this gap fully. PCR targeting of the SeM gene successfully identified S. equi in 49 (83.1%) out of the 59 isolates. These results were consistent with previous studies by Alwan (Alwan, 2014) in Iraq, who identified S. equi 24/30 (80%) isolates using the SeM gene with Real-Time Polymerase Chain Reaction (Real-Time PCR), and Tartor (Tartor et al., 2020) in Egypt, who successfully identified S. equi 92/100 (92%). Mahmood (Mahmood et al., 2014) identified 51/101 (50.49%) positive samples from equine clinical samples using the SeM gene in Iraq. In Colombia, Morales (Jaramillo-Morales et al., 2022) reported a (13.5%) prevalence of S. equi in guttural pouch samples. These variations in detection rates may reflect differences in sampling methods, populations, and diagnostic approaches across studies. Analysis of our SeM gene sequences revealed four polymorphisms in comparison to the reference S. equi 4047 strain. These polymorphisms led to amino acid changes and may influence the virulence and immunogenicity of S. equi. The exact match with an Iranian strain (OL332314.1) (Moghaddam et al., 2023) suggests a potential link in terms of epidemiology, highlighting the need for regional surveillance and comparative genomic studies. This study provides the first report of SeM allele typing in Iraq, identifying the SeM-97 allele in S. equi isolates. The SeM-97 allele was previously reported by (Moghaddam et al., 2023) in Iran, highlighting potential epidemiological links between the regions. The identification of SeM alleles aids in differentiating strangles outbreaks and understanding the epidemiology of the disease at regional and national levels (Ivens et al., 2011; Parkinson et al., 2011). SeM typing in other regions emphasizes its value in understanding strain movement and epidemiology. In Egypt, SeM typing revealed novel alleles (SeM-139 and SeM-141) linked to strains from donkeys in China and others (SeM-140, SeM- 199, and SeM-7) associated with European isolates (Tartor et al., 2020). Similarly, in Thailand, the identification of SeM-48, previously reported in UK outbreaks, suggests the importation of infected horses as a potential source of transmission (Tonpitak et al., 2016). These studies demonstrate how SeM allele data can be used to trace cross-regional strain connections and inform control measures. ConclusionThis study provides critical molecular and sequencing insights into the S. equi strains responsible for strangles in Baghdad, Iraq. The detection of the SeM- 97 allele, reported for the first time in Iraq, highlights the utility of molecular diagnostics in identifying and characterizing strains associated with outbreaks. Furthermore, these results support the hypothesis of regional strain connections due to unsupervised horse importation. These findings establish a foundation for future epidemiological research and emphasize the need for targeted biosecurity measures and control strategies to mitigate strangles in Iraq. AcknowledgmentsThe authors thank the College of Veterinary Medicine, University of Baghdad, for providing the necessary facilities and support to conduct this study. We also extend our gratitude to the laboratory staff for their invaluable assistance. Conflicts of interestThe authors declare no conflicts of interest. FundingThis research received no specific grant from funding agencies in the public, commercial, or not-for-profit sectors. Authors’ contributionsSaif Aldeen Kamal Abdul-Latif: Conceptualization, sample collection, data analysis, methodology, molecular work, manuscript drafting, and final review. Afaf Abdulrahman Yousif: Supervision and critical manuscript review. Both authors reviewed and approved the final manuscript. Data availabilityThe SeM gene sequences analyzed during this study have been submitted to the NCBI GenBank database under the accession numbers PP726110.1, PP738564.1, PP738563.1, PP738562.1, PP738561.1, PP738560.1, PP738559.1, PP738558.1, PP738557.1. ReferencesAl-Robaiee, I.A., Hassan, W.S. and Al-Farwachi, M.I. 2023. Seroprevalence of strangles in horses and donkeys in Mosul city, Iraq. J. Adv. Vet. Res. 14(1), 158–160. Alwan, J.A. 2014. Used of real-time PCR technique for detection of Streptococcus equi from horses infected by strangles in AL-Diwanyia Provence, Iraq. Int. J. Adv. Res. 2(4), 208–213. Anzai, T., Timoney, J.F., Kuwamoto, Y., Fujita, Y., Wada, R. and Inoue, T. 1999. In vivo pathogenicity and resistance to phagocytosis of Streptococcus equi strains with different levels of capsule expression. Vet. Microbiol. 67(4), 277–286. Arafa, A., El-Razik, K., Kandil, M. and Abdalhamed, A. 2024. Phenotypic and genotypic characterization of Escherichia coli isolates recovered from equines with respiratory problems in Egypt. Int. J. Vet. Sci. 14(2), 260–267. Arafa, A.A., Hedia, R.H., Ata, N.S. and Ibrahim, E.S. 2021. Vancomycin resistant Streptococcus equi subsp. equi isolated from equines suffering from respiratory manifestation in Egypt. Vet. World 14(7), 1808–1814. Artiushin, S.C., Timoney, J.F., Sheoran, A.S. and Muthupalani, S.K. 2002. Characterization and immunogenicity of pyrogenic mitogens SePE-H and SePE-I of Streptococcus equi. Microb. Pathog. 32(2), 71–85. Baverud, V., Johansson, S.K. and Aspan, A. 2007. Real-time PCR for detection and differentiation of Streptococcus equi subsp. equi and Streptococcus equi subsp. zooepidemicus. Vet. Microbiol. 124(3–4), 219–229. Bekele, D., Dessalegn, B., Tadesse, B. and Abey, S.L. 2024. Streptococcus equi subspecies equi from strangles suspected equines: molecular detection, antibiogram profiles and risk factors. BMC Vet. Res. 20(1), 377. Boschwitz, J.S. and Timoney, J.F. 1994. Inhibition of C3 deposition on Streptococcus equi subsp. equi by M protein: a mechanism for survival in equine blood. Infect Immun. 62(8), 3515–3520. Boyle, A.G. 2016. Strangles and its complications. Equine Vet. Educ. 29(3), 149–157. Boyle, A.G., Sweeney, C.R., Kristula, M., Boston, R. and Smith, G. 2009. Factors associated with likelihood of horses having a high serum Streptococcus equi SeM-specific antibody titer. J. Am. Vet. Med. Assoc. 235(8), 973–977. Dong, J., Gao, N., Waller, A.S., Cook, F.R., Fan, S., Yuan, D., Du, Y., Li, F., Norimine, J. and Zhu, W. 2019. An outbreak of strangles associated with a novel genotype of Streptococcus equi subspecies equi in donkeys in China during 2018. Equine Vet. J. 51(6), 743–748. Duran, M.C. and Goehring, L.S. 2021. Equine strangles: an update on disease control and prevention. Austral. J. Vet. Sci. 53(1), 23–31. Durham, A.E. and Kemp-Symonds, J. 2021. Failure of serological testing for antigens A and C of Streptococcus equi subspecies equi to identify guttural pouch carriers. Equine Vet. J. 53(1), 38–43. Edgar, R.C. 2004. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32(5), 1792–1797. Gronbaek, L.M., Angen, O., Vigre, H. and Olsen, S. N. 2006. Evaluation of a nested PCR test and bacterial culture of swabs from the nasal passages and from abscesses in relation to diagnosis of Streptococcus equi infection (strangles). Equine Vet. J. 38(1), 59–63. Guindon, S., Dufayard, J.F., Lefort, V., Anisimova, M., Hordijk, W. and Gascuel, O. 2010. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59(3), 307–321. Holden, M.T., Heather, Z., Paillot, R., Steward, K.F., Webb, K., Ainslie, F., Jourdan, T., Bason, N.C., Holroyd, N.E., Mungall, K., Quail, M.A., Sanders, M., Simmonds, M., Willey, D., Brooks, K., Aanensen, D.M., Spratt, B.G., Jolley, K.A., Maiden, M.C., Kehoe, M., Chanter, N., Bentley, S.D., Robinson, C., Maskell, D.J., Parkhill, J. and Waller, A.S. 2009. Genomic evidence for the evolution of Streptococcus equi: host restriction, increased virulence, and genetic exchange with human pathogens. PLoS Pathog. 5(3), e1000346. Ivens, P.A., Matthews, D., Webb, K., Newton, J.R., Steward, K., Waller, A.S., Robinson, C. and Slater, J.D. 2011. Molecular characterisation of ‘strangles’ outbreaks in the UK: the use of M-protein typing of Streptococcus equi ssp. equi. Equine Vet. J. 43(3), 359–364. Jaramillo-Morales, C., Gomez, D.E., Renaud, D. and Arroyo, L.G. 2022. Streptococcus equi culture prevalence, associated risk factors and antimicrobial susceptibility in a horse population from Colombia. J. Equine Vet. Sci. 111, 103890. Jaramillo-Morales, C., James, K., Barnum, S., Vaala, W., Chappell, D.E., Schneider, C., Craig, B., Bain, F., Barnett, D.C., Gaughan, E. and Pusterla, N. 2023. Voluntary biosurveillance of Streptococcus equi subsp. equi in nasal secretions of 9409 equids with upper airway infection in the USA. Vet. Sci. 10(2), 78. Jawad, A.N.M. 2000. Clinical study on strangles in Iraqi horses. Iraqi J. Vet. Med. 24(2), 1–4. Kaisium, M.M. and Al-Judi, A.M.H. 2007. Preparation of live attenuated vaccine against strangles. Iraqi J. Vet. Med. 31(1), 50–54. Kasuya, K., Tanaka, N., Oshima, F., Fujisawa, N., Saito, M., Tagami, K., Niwa, H. and Sasai, K. 2019. Genetic analysis of Streptococcus equi subsp. equi isolated from horses imported into Japan. J. Vet. Med. Sci. 81(6), 924–927. Katoh, K. and Standley, D.M. 2013. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30(4), 772–780. Kelly, C., Bugg, M., Robinson, C., Mitchell, Z., Davis-Poynter, N., Newton, J.R., Jolley, K.A., Maiden, M.C. and Waller, A.S. 2006. Sequence variation of the SeM gene of Streptococcus equi allows discrimination of the source of strangles outbreaks. J. Clin. Microbiol. 44(2), 480–486. Letunic, I. and Bork, P. 2024. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52(W1), W78–W82. Libardoni, F., Vielmo, A., Farias, L., Matter, L.B., Pötter, L., Spilki, F.R. and de Vargas, A.C. 2013. Diversity of seM in Streptococcus equi subsp. equi isolated from strangles outbreaks. Vet. Microbiol. 162(2–4), 663–669. Lindahl, S., Soderlund, R., Frosth, S., Pringle, J., Baverud, V. and Aspan, A. 2011. Tracing outbreaks of Streptococcus equi infection (strangles) in horses using sequence variation in the seM gene and pulsed-field gel electrophoresis. Vet. Microbiol. 153(1–2), 144–149. Liu, Y. 2019. Identification of a novel genotype of Streptococcus equi subspecies equi in a donkey suffering from strangles. Pak. Vet. J. 39(4), 609–611. Mahmood, S.H., Al-Judi, A.M. and Mosstafa, H.K. 2014. Molecular identification of Streptococcus equi subspecies equi in horses. Iraqi J. Vet. Med. 38(2), 1–8. McGlennon, A., Waller, A., Verheyen, K., Slater, J., Grewar, J., Aanensen, D. and Newton, R. 2021. Surveillance of strangles in UK horses between 2015 and 2019 based on laboratory detection of Streptococcus equi. Vet. Rec. 189(12), e948. Meehan, M., Muldowney, D.A., Watkins, N.J. and Owen, P. 2000. Localization and characterization of the ligand-binding domain of the fibrinogen-binding protein (FgBP) of Streptococcus equi subsp. equi. Microbiology (Reading). 146(Pt 5), 1187–1194. Moghaddam, S., Lotfollahzadeh, S., Salehi, T.Z., Hassanpour, A., Manesh, H.T. and Tamai, I.A. 2023. Molecular and sequencing study and identification of novel SeM-type in beta-hemolytic streptococci involving the upper respiratory tract in Iran. BMC Vet. Res. 19(1), 210. Moloney, E., Kavanagh, K.S., Buckley, T.C. and Cooney, J.C. 2013. Lineages of Streptococcus equi ssp. equi in the Irish equine industry. Ir. Vet. J. 66(1), 10. Nei, M. and Kumar, S. 2000. Molecular evolution and phylogenetics. Oxford, UK: Oxford University Press. Newton, J.R., Wood, J.L., Dunn, K.A., DeBrauwere, M.N. and Chanter, N. 1997. Naturally occurring persistent and asymptomatic infection of the guttural pouches of horses with Streptococcus equi. Vet. Rec. 140(4), 84–90. Parkinson, N.J., Robin, C., Newton, J.R., Slater, J. and Waller, A.S. 2011. Molecular epidemiology of strangles outbreaks in the UK during 2010. Vet. Rec. 168(25), 666. Patty, O.A. and Cursons, R.T. 2014. The molecular identification of Streptococcus equi subsp. equi strains isolated within New Zealand. N Z Vet. J. 62(2), 63–67. Proft, T., Webb, P.D., Handley, V. and Fraser, J.D. 2003. Two novel superantigens found in both group A and group C Streptococcus. Infect Immun. 71(3), 1361–1369. Quinn, P.J., Markey, B.K., Leonard, F.C., FitzPatrick, E.S., Fanning, S. and Hartigan, P.J. 2011. Veterinary microbiology and microbial disease. Hoboken, NJ: Wiley-Blackwell. Robinson, C., Steward, K.F., Potts, N., Barker, C., Hammond, T.A., Pierce, K., Gunnarsson, E., Svansson, V., Slater, J., Newton, J.R. and Waller, A.S. 2013. Combining two serological assays optimises sensitivity and specificity for the identification of Streptococcus equi subsp. equi exposure. Vet. J. 197(2), 188–191. Rotinsulu, DA., Ewers, C., Kerner, K., Amrozi, A., Soejoedono, R.D., Semmler, T. and Bauerfeind, R. 2023. Molecular features and antimicrobial susceptibilities of Streptococcus equi ssp. equi isolates from strangles cases in Indonesia. Vet. Sci. 10(1), 49. Sweeney, C.R., Benson, C.E., Whitlock, R.H., Meirs, D.A., Barningham, S.O., Whitehead, S.C. and Cohen, D. 1989. Description of an epizootic and persistence of Streptococcus equi infections in horses. J. Am. Vet. Med. Assoc. 194(9), 1281–1286. Sweeney, C.R., Timoney, J.F., Newton, J.R. and Hines, M.T. 2005. Streptococcus equi infections in horses: guidelines for treatment, control, and prevention of strangles. J. Vet. Intern. Med. 19(1), 123–134. Tamura, K., Stecher, G. and Kumar, S. 2021. MEGA11: molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38(7), 3022–3027. Tartor, Y.H., El-Naenaeey, E.Y., Gharieb, N.M., Ali, W.S. and Ammar, A.M. 2020. Novel Streptococcus equi strains causing strangles outbreaks in Arabian horses in Egypt. Transbound Emerg. Dis. 67(6), 2455–2466. Timoney, J.F. and Artiushin, S.C. 1997. Detection of Streptococcus equi in equine nasal swabs and washes by DNA amplification. Vet. Rec. 141(17), 446–447. Timoney, J.F., Suther, P., Velineni, S. and Artiushin, S.C. 2014. The antiphagocytic activity of SeM of Streptococcus equi requires capsule. J. Equine Sci. 25(2), 53–56. Tonpitak, W., Sornklien, C. and Wutthiwithayaphong, S. 2016. Characterization of a Streptococcus equi ssp. equi isolate from a strangles outbreak in Thailand. J. Equine Vet. Sci. 38, 30–32. Whelchel, D.D. and Chaffin, M.K. 2009. Sequelae and complications of Streptococcus equi subspecies equi infections in the horse. Equine Vet. Educ. 21(3), 135–141. Zu, H., Sun, R., Li, J., Guo, X., Wang, M., Guo, W. and Wang, X. 2024. Development of a real-time recombinase-aided amplification method for the rapid detection of Streptococcus equi subsp. equi. Microorganisms 12(4), 777. | ||
| How to Cite this Article |
| Pubmed Style Abdul-latif SAK, Yousif AA. Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Vet. J.. 2025; 15(2): 731-737. doi:10.5455/OVJ.2025.v15.i2.22 Web Style Abdul-latif SAK, Yousif AA. Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. https://www.openveterinaryjournal.com/?mno=225994 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i2.22 AMA (American Medical Association) Style Abdul-latif SAK, Yousif AA. Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Vet. J.. 2025; 15(2): 731-737. doi:10.5455/OVJ.2025.v15.i2.22 Vancouver/ICMJE Style Abdul-latif SAK, Yousif AA. Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Vet. J.. (2025), [cited January 25, 2026]; 15(2): 731-737. doi:10.5455/OVJ.2025.v15.i2.22 Harvard Style Abdul-latif, S. A. K. & Yousif, . A. A. (2025) Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Vet. J., 15 (2), 731-737. doi:10.5455/OVJ.2025.v15.i2.22 Turabian Style Abdul-latif, Saif Aldeen Kamal, and Afaf Abdulrahman Yousif. 2025. Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Veterinary Journal, 15 (2), 731-737. doi:10.5455/OVJ.2025.v15.i2.22 Chicago Style Abdul-latif, Saif Aldeen Kamal, and Afaf Abdulrahman Yousif. "Molecular study of Streptococcus equi isolated from horses with strangles in Iraq." Open Veterinary Journal 15 (2025), 731-737. doi:10.5455/OVJ.2025.v15.i2.22 MLA (The Modern Language Association) Style Abdul-latif, Saif Aldeen Kamal, and Afaf Abdulrahman Yousif. "Molecular study of Streptococcus equi isolated from horses with strangles in Iraq." Open Veterinary Journal 15.2 (2025), 731-737. Print. doi:10.5455/OVJ.2025.v15.i2.22 APA (American Psychological Association) Style Abdul-latif, S. A. K. & Yousif, . A. A. (2025) Molecular study of Streptococcus equi isolated from horses with strangles in Iraq. Open Veterinary Journal, 15 (2), 731-737. doi:10.5455/OVJ.2025.v15.i2.22 |