| Research Article | ||
Open Vet. J.. 2025; 15(2): 765-773 Open Veterinary Journal, 2025), Vol. 15(2): 765-773 Research Article Molecular detection of coronavirus in camelids and bovines using real-time quantitative polymerase chain reaction in Wasit Province, IraqIbrahim Hasan Madhloom1, Rasha Munther Othman1 and Hussein Ali Mohammed Al-Bayati21Department of Microbiology, College of Veterinary Medicine, University of Basrah, Basrah, Iraq 2Department of Microbiology, College of Science, University of Wasit, Kut, Iraq *Corresponding Author: I. H. Madhloom. Department of Microbiology, College of Veterinary Medicine, University of Basrah, Basrah, Iraq. Email: ibrahim1987hasan [at] gmail.com Submitted: 28/10/2024 Accepted: 08/01/2025 Published: 28/02/2025 © 2025 Open Veterinary Journal
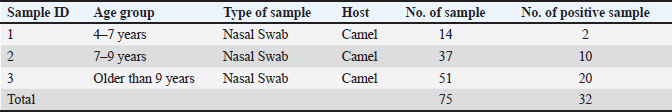
AbstractBackground: Coronaviruses (CoVs) are a diverse group of RNA viruses that cause respiratory and gastrointestinal diseases in humans and animals. Over the past two decades, outbreaks of Middle East Respiratory Syndrome Coronavirus (MERS-CoV) and bovine coronavirus (BCoV) have affected animal populations, especially in regions with close animal-human interactions, such as the Arabian Peninsula and Iraq. Given the potential for zoonotic transmission, understanding the prevalence and spread of CoVs among livestock is essential for managing potential risks to animal and human health. Aim: This study aimed to investigate the prevalence of MERS-CoV in camels and BCoV in bovines within the Wasit Governorate of Iraq to assess the infection rates and potential interspecies transmission risks. Methods: One hundred and fifty nasal swab samples (75 from camels and 75 from bovines) were collected between November 2022 and April 2023. The samples were analyzed for the presence of MERS-CoV and BCoV using real-time quantitative reverse transcription PCR (qRT-PCR) targeting the nucleocapsid (N) gene for each virus. Standard procedures for RNA extraction were followed, and qRT–PCR assays were conducted using specific primers to ensure high sensitivity and specificity. Results: MERS-CoV was present in (42%) of the camel samples, whereas BCoV was detected in (34%) of the bovine samples. Statistical analysis indicated a significant difference (p < 0.05) in infection rates between camels and bovines, with a higher prevalence observed in camels. The clinical signs observed in infected camels included fever, nasal discharge, and appetite loss, whereas infected bovines exhibited symptoms such as diarrhea and respiratory distress. Conclusion: The high prevalence of MERS-CoV and BCoV in camels and bovines in the Wasit region indicates a substantial risk for the continued spread of these viruses within animal populations. These findings underscore the importance of surveillance and biosecurity measures to control the spread of coronavirus among livestock, potentially reducing zoonotic transmission risks. Further research is required to understand the transmission dynamics of CoVs in mixed livestock farming systems. Keywords: Coronaviruses, MERS-CoV, BCoV, RNA viruses, qRT-PCR, zoonotic transmission. IntroductionCoronaviruses (CoVs) cause disease in humans and animals (Saied et al., 2021). In the past 20 years, three global outbreaks have occurred: Coronavirus Disease-2019 in Wuhan, China; Middle East Respiratory Syndrome Coronavirus (MERS-CoV) in Saudi Arabia; and Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) in China (Colina et al., 2021). Dromedaries are widely recognized as natural hosts of MERS-CoV, which can infect humans and domestic animals (Kandeil et al., 2019). While dromedary camels represent the virus reservoir and a means of human transmission, bats are believed to have been the virus’s original host (Al-Shomrani et al., 2020). MERS-CoV infection in humans may persist due to intimate interactions between infected dromedaries and humans, facilitating zoonotic transmission (De Wit et al., 2016). Bovine coronavirus (BCoV) causes neonatal calf diarrhea in newborn calves, winter dysentery in adult bovines, and respiratory tract infections in all ages (Johnson and Pendell, 2017). Bovine respiratory coronavirus (BRCoV) is a strain of coronavirus isolated from the respiratory tract, whereas bovine enteric coronavirus (BECoV) is a strain of coronavirus isolated from the intestinal tract. The BECoV strains are responsible for calf diarrhea, and winter dysentery are further classified as BECoV-CD and BECoV-WD, respectively (Boileau and Kapil, 2010). CoVs belong to the Nidovirales order and the subfamily Coronavirinae of the Coronaviridae family (Lai et al., 1994). The Coronavirinae subfamily is divided into four genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus (α-, β-, γ-, and δ-CoV) based on genomic structure and evolutionary relationships. CoVs from mammals are generally included in α-CoVs and β-CoVs, while CoVs from birds are mainly included in γ-CoVs and δ-CoVs-CoVs (Li et al., 2022). The α-CoVs and β-CoVs are of greater interest due to their ability to penetrate animal-human barriers and cause significant human infections (Coleman and Frieman, 2014). CoV chimeric are enclosed viruses with the most giant known RNA virus genomes, with a single-stranded, positive-sense RNA genome ranging from 26 to 32 kilobases (Weiss and Navas-Martin, 2005). Electron microscopy revealed spike projections from CoV virions resembling a crown, or corona in Latin, which is the term used to describe the virus (Pal et al., 2020). All CoVs have identical genomic structures and expressions, with structural proteins, such as spikes, envelopes, membranes, and nucleocapsid, originating from the open reading frame 1a/b-encoded nsp1 through nsp16 (Su et al., 2016). The hemagglutinin-esterase accessory protein, which is found exclusively in certain beta-CoVs, forms a smaller layer of projections within the betacoronavirus genome (Masters, 2006). The viral envelope’s helical capsid structure, formed by nucleocapsid protein binding with genomic RNA, creates spikes in some CoV virions, while transmembrane proteins “M” and “E” aid virus assembly (Weiss and Leibowitz, 2011). Real-time quantitative polymerase chain reaction (RT-qPCR) is frequently utilized for detecting gene expression levels and diagnosing acute respiratory viral infections quickly (Corman et al., 2020). Conventional real-time RT-PCR assays require large laboratory instruments and take approximately 2 hours to amplification (Shirato et al., 2020). MERS-COVs and BCoVs can be diagnosed in the laboratory by detecting their genes in clinical samples from diseased animals (Decaro et al., 2008; Shirato et al., 2020). The standard method for diagnosing MERS-CoV is RT-PCR (Al Johani and Hajeer, 2016). Cattle rectal and respiratory samples can be used to detect and identify BCoV and BCoV-like RNA using RT-PCR–based techniques (Erles et al., 2003; Takeuchi et al., 2006). The Iraqi government and official institutions have shown limited or no interest in the surveillance or vaccination efforts for MERS-CoV and BCoV. Currently, there are no established national programs targeting these viruses. Efforts in this field are mainly driven by individual researchers, focusing on specific aspects, such as the partial genetic or immunological characterization of CoVs in camels and cattle. This highlights the need for greater institutional involvement to address potential public health and veterinary concerns. In general, viral diseases in veterinary medicine are a significant threat to animal health and can cause considerable economic losses. Recent studies conducted in southern Iraq highlight the importance of understanding these diseases from various perspectives (Thwiny et al., 2015; Thwiny, 2016; Thwiny et al., 2018; Al-Mubarak et al., 2022; Al-Mubarak et al., 2023). Together, these studies emphasize the need for comprehensive approaches to manage viral diseases in veterinary medicine in southern Iraq. This study aimed to investigate the presence of CoVs in camel and bovine samples through real-time quantitative polymerase chain reaction (qPCR) analysis, with the objective of elucidating viral prevalence and identifying potential risk factors associated with transmission within these animal populations. Ethical approval and animal welfare statementThis study was conducted according to ethical guidelines and was approved by the Scientific Committee of the College of Veterinary Medicine, University of Basrah (Approval Number:39/2024). All procedures involving animals complied with institutional and international standards for animal welfare. Efforts were made to minimize animal suffering and ensure proper care during the study. Materials and MethodsSample collectionThe methods of sample collection in this study were as follows: Nasal swabs were collected from 75 camels to detect viral shedding from the respiratory tract. In addition, nasal and rectal swabs were collected from 75 bovines to detect respiratory and gastrointestinal shedding of the virus. We obtained samples from animals across different age groups in Wasit Province, Iraq, between November 2022 and April 2023. Sample collection followed standard protocols to minimize animal stress and discomfort, ensuring ethical considerations were adhered to during the process. We collected swabs from all hosts using a virus sampling kit (Jiangsu, China). We stored the samples on ice during transportation to the microbiology laboratory in the College of Medicine, University of Wasit, where they were stored at −80°C until they were thawed and used for RNA extraction and CoV testing. We have automated RNA extractionCoronavirus RNA was extracted automatically using a nucleic acid extraction kit (magnetic bead method) from Chongqing, China. We added 15 µl of proteinase K to mix wells A1–H1 and A7–H7, followed by 200 µl of viral transport medium sample, following the manufacturer’s guidelines. The plates were placed in the EXM3000 isolation system, and the extraction process was allowed to run for 9 minutes. qRT-PCRFollowing the manufacturer’s instructions, a GoTaq®1-Step RT-qPCR System Promega kit (Wisconsin, USA) was used for a one-step qRT–PCR assay. The final GoTaq® 1-Step RT-qPCR reaction mixture for qRT-PCR consisted of 10 μl of GoTaq® qPCR Master Mix 2×, 2 μl of forward primer 10×, 2 μl of reverse primer 10×, 0.4 μl of GoScript™ RT Mix for 1-Step RT-qPCR, 4 μl of extracted RNA, and up to 20 µl of nuclease-free water. The mixture underwent one reverse transcription phase for 15 minutes at 38°C, an initial denaturation step of 10 minutes at 95°C, and 40 cycles of 10 seconds at 95°C, 30 seconds at 58°C, and 30 seconds at 72°C. PrimersFor qRT-PCR, we used oligonucleotide primers targeting two distinct genomic sites: the gene fragment specific to MERS-CoVs in camels. Based on the nucleocapsid gene’s start codon, the primer sequence was as follows: the upstream primers 5’-TGCAAGCTTTTGGTCTTCGC-3’ and 5’-AGCAAGCTCAGCAATTTGGG-3’ are used to synthesize the desired product (Al Salihi, 2017). The N gene fragment for the Mebus strain in bovines was sequenced using the upstream primer 5’-GCAATCCAGTAGTAGAGCGT-3’ and the downstream primer 5’-CTTAGTGGCATCCTTGCCAA-3’ are both essential components in the qRT-PCR synthesis process (Hasoksuz et al., 2002). Statistical analysesThe data were analyzed using SPSS software and Chi-square tests to establish the relationship between MERS-CoV in camels and BCoV in bovines and calculate the percentages of infected animals. ResultsClinical cases were previously diagnosed using clinical findings from diseased animals such as camels and bovines (Table 3 and Fig. 1). Clinical findings revealed fever, ocular, and nasal discharge, and appetite loss in camels aged 4–14 years (Table 1 and Fig. 3). During data collection and case history gathering from camel owners, it was clarified that there was an unlimited number of fetuses with gray eyes at different stages of gestation in pregnant females. Additionally, it was noted that the camel mortality rate during the study period was (3%) during the study period. The disease in bovines manifests as soft, runny, and weak diarrhea in calves aged 7–21 days, green watery diarrhea in calves aged 3–9 months, and persistent cough and nasal discharge in cows aged 4 years (Table 2 and Fig. 2). Table 1. Sample collection overview: Nasal swabs from camels by age group.
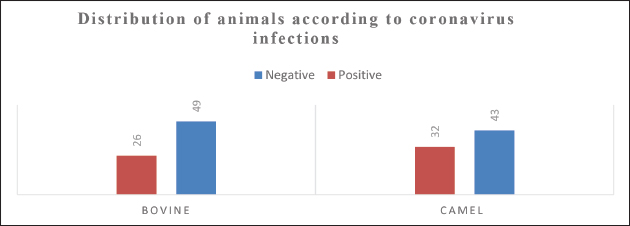
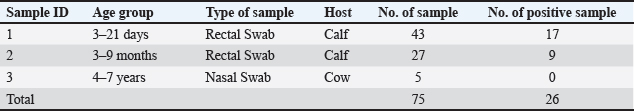
Figure 1. The study examines the distribution of animals based on infection by BCoVs and MERS-CoVs. Table 2. Sample collection overview: Rectal and nasal swabs from cattle by age group.
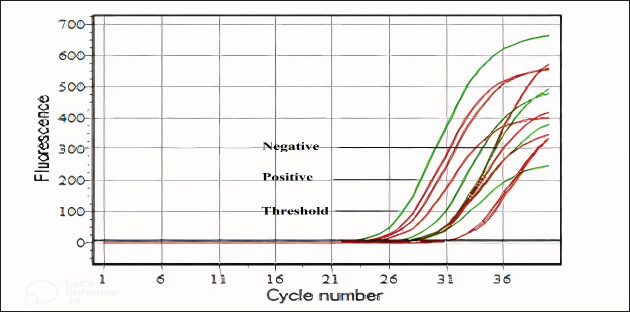
Figure 2. Amplification plot curves in bovine indicate a correlation between the fluorescence of the Fluorescein Amidite (FAM) channel and cycle number. The threshold, positive, and negative samples were tested using the RTqPCR device. The threshold based on a positive sample ranged from 28 to 31 cycles. Table 3. Animal distribution based on coronavirus infection rates.
Figure 3. The amplification plot curves in camles indicate a correlation between the fluorescence of the FAM channel and the cycle number. The threshold, positive, and negative samples were tested using an RT-qPCR device. The threshold based on a positive sample ranged from 27 to 31 cycles. DiscussionThis unique study in Iraq aimed to explore camels and bovines as hosts of the coronavirus, which causes various diseases, such as respiratory and digestive diseases in camels. The new study’s results and previous research are entirely out of the ordinary; the study revealed a substantial increase in the coronavirus infection rate in camels and bovines (Alrodhan, 2017; Mansour et al., 2013). This phenomenon dates back to the period accompanying our study of the advanced strategic shifts that characterize the coronavirus, which usually increases its spread and the strength of its infection. However, some coronavirus genera pursue a niche evolutionary strategy to infect a specific host and bypass others, as occurs in gammacoronavirus, which only infects birds. According to recent molecular and serological evidence, the dromedary camel is the host of MERS-CoV. Before 2012, MERS-CoV or MERS-like CoV was circulating among dromedaries in Africa and the Arabian Peninsula (Reusken et al., 2014). RT-PCR confirmed the presence of MERS-CoV in nasal samples, live and lung tissue samples, and carcasses from dromedary camels in various Arabian Peninsula locations as evidence (Khalafalla et al., 2015). As previously mentioned, MERS-CoV was confirmed in camel samples by screening them with MERS-CoV-specific RT-PCR targeting the N gene (Corman et al., 2012). Research on camels has shown that MERS-CoVs are prevalent in various countries (Alagaili et al., 2014; Chu et al., 2014; Hemida et al., 2014; Wernery et al., 2015). The first Iraqi study using RT-PCR to detect and characterize MERS-CoV in dromedaries and humans found MERS-CoV present in (15%) and (5%) of the population, respectively (Al Salihi, 2017). Previous research has focused on genetic characterization and the prevalence of MERS-CoVs in Qatar (1.9%), UAE (1.6%), Egypt (15.4%), Jordan (93.3%), Kenya (0.9%), and Pakistan (2.8%) (Raj et al., 2014; Yusof et al., 2015; Ali et al., 2017; van Doremalen et al., 2017; Ommeh et al., 2018; Zohaib et al., 2018). This study found that (42.6%) of camel nasal swabs were detected by qRT-PCR, contradicting previous findings suggesting different MERS-CoV infection rates between 2014 and 2018, with the highest rate in Jordan in 2017. The paucity of studies and the absence of prospective surveillance of MERS-CoV in camels in Iraq contribute to our imperfect understanding of the virus’s epidemiology and the risk factors associated with its spread. However, when comparing the results of this study in Iraq with earlier data (Al Salihi, 2017), it is evident that the prevalence of MERS-CoV has increased over time. Owners of camels in Badra, Iraq, reported no disease, but during grazing in the Middle Euphrates, they experienced high fever, abortion, and stillbirth. A previous study revealed that camelid infections have a high transmission potential and are characterized by widespread viral shedding and upper respiratory tract replication (Te et al., 2022). Based on the findings that MERS-CoV is typically shed from diseased camels through nasal discharge and remains viable and environmentally stable on various surfaces under different temperatures and humidity levels, particularly at lower temperatures and humidity (Van Doremalen, 2013), and considering the practices of camel owners who rotate their herds in specific grazing areas. We hypothesize that virus shedding from camels, such as through nasal discharge, contributes to the environmental persistence of MERS-CoV on grass or feed. The virus remains viable, stable, and infectious on contaminated grass, facilitating its transmission. This transmission occurs through the ingesting or inhalation of contaminated grass between camel herds, demonstrating the virus’s environmental stability and its role in facilitating interherd transmission. MERS-CoVs spread among dromedaries, and their strains exhibited high genetic similarity to human isolates (Omrani et al., 2015). Moreover, sheep, goats, and donkeys grown close to camels tested positive for MERS-CoV (Kandeil et al., 2019). Recent studies have shown that dromedaries play crucial roles in the spread of MERS-CoV at the animal-human interface (Haagmans et al., 2014). Dromedaries can transmit MERS-COVs to humans through direct contact with mucous and nasal secretion or by consuming meat or raw milk (Gossner et al., 2016; Mirkena et al., 2018). MERS-CoV RNA was detected in milk from milking animals with camels of different origins, as confirmed by qRT-PCR (Reusken et al., 2016). Identifying MERS-CoV infection in dromedaries is challenging because of its asymptomatic nature, but experimental cases show tracheitis, bronchitis, rhinitis, nasal discharge, and significant virus release (Haverkamp et al., 2018). From 2010 to 2019, multiple countries confirmed positive BCoV infections in calf feces following the virus’s molecular detection using qRT–PCR (Zhu et al., 2022). Including Iraq (6.57%) (Mansour et al., 2013) and Vietnam (6.9%) (Shin et al., 2019), China (18.9.0%) (Keha et al., 2019), Thailand (12%) (Singasa et al., 2017), India (9.38%) (Kumar et al., 2013), Korea (6.2%) (Lee et al., 2019), and Korea (5.9%) (Kim et al., 2022). In 2002–2003, the prevalence of BCoVs was (33.0%) in Korea and remained between (5.4%) and (15.6%) until 2021 (Jeong et al., 2005). This study indicates an increase in the prevalence of BCoVs in Iraq, but it suggests that continuous and potentially harmful infections in bovine farms may persist. This study revealed a (34.6%) prevalence of BCoVs in diarrheal calves on bovine farms, highlighting the need for ongoing epidemiological surveillance to prevent future outbreaks. This study suggests that the increase in BCoV infections compared with the Iraqi 2013 study and other international studies may be attributable to various factors. First, the studies show a time gap, suggesting that BCoVs developed an evolution strategy to increase their spread during grazing or feedlot bovines’ fields. The second reason is the implementation of preventive programs in Western and Asian countries, such as BCoV vaccines, which significantly decrease the incidence of infections. Research indicates that the virus persists in subclinically infected adult cattle (Park et al., 2007; Toftaker et al., 2017). BCoV can be shed for up to 14 days through high concentrations of virus particles in respiratory and gastrointestinal secretions (1 billion per milliliter of faeces) (Kapil et al., 1990). Moreover, the high density of animals appears to be the primary risk factor for BCoVs (Boileau and Kapil, 2010). Diarrhea in calves is primarily caused by the BCoV virus during the first month of life (Brandão et al., 2006; Ammar et al., 2014). A study suggests that (10%–30%) of newborn diarrhea cases may be caused by BCoV (Alfieri et al., 2018). BCoV is linked to various diseases, including respiratory infections like shipping fever in young cattle and enzootic pneumonia in calves (Saif, 2010). Environmental studies have shown that animal CoVs remain infectious in water and sewage for up to a year, depending on the temperature and humidity (Mullis et al., 2012). Previous studies have shown that the temperature of the surrounding environment significantly influences the stability of CoVs (Siddell et al., 1983; Tennant et al., 1994). CoVs are inactivated at 56°C for 10–15 minutes, 37°C for a few days and 4°C for a few months. In contrast, CoVs can survive at −60°C for several years without losing their ability to spread (Andries et al., 2010; McIntosh et al., 1974; Siddell et al., 1983). Recent environmental survival research on two animals, CoVs, revealed that they can survive for weeks at 27°C in water and sewage and up to a year at 4°C (Casanova et al., 2009). ConclusionBetacoronavirus infects camels and bovines and causes diseases in the respiratory and digestive systems of infected animals, leading to their owners incurring significant economic losses. The study revealed that camels have high temperatures, runny noses, and eyes, while bovines have watery diarrhea and sometimes bleeding. According to our findings, the coronavirus spreads quickly on inanimate surfaces and grass, as it remains on these surfaces for long periods. It is recommended that RT-qPCR technology be applied for accurate diagnosis of coronavirus in camels and bovines, emphasizing the importance of managing, following up on, and monitoring the spread of this deadly virus in Iraq. AcknowledgmentsWe express our sincere gratitude to Captain Haider Al-Badiri from the Iraqi Camel Federation for his invaluable assistance in collecting camel samples. We would like to extend our heartfelt thanks to Dr Khalid Sajit from the Veterinary Hospital in Wasit, Veterinary Directorate, Ministry of Agriculture, Iraq, for his significant support in collecting cattle samples. Their contributions are crucial to the success of this research. Conflict of interestThe authors affirm that there are no conflicts of interest, whether financial, personal, or related to any other relationships, that may affect the integrity of the research presented in this manuscript. Author contributionsThe specific contributions of each author are indicated with initials, i.e. Conception and design of the study: RM and HA. Acquisition of data: IH. Analysis and/or interpretation of the data: HA and IH. Drafting of the manuscript: RM and HA. Critical review/revision: RM and HA. ReferencesAl Johani, S. and Hajeer, A.H. 2016. MERS-CoV diagnosis: an update. J. Infect. Public Health. 9(3), 216–219. Al Salihi, S.F. 2017. Phylogenetic analysis of MERSCoV in human and camels in Iraq. AL-Qadisiyah Med. J. 13(23), 150–159. Alagaili, A.N., Briese, T., Mishra, N., Kapoor, V., Sameroff, S.C., de Wit, E., Munster, V.J., Hensley, L.E., Zalmout, I.S., Kapoor, A., Epstein, J.H., Karesh, W.B., Daszak, P., Mohammed, O.B. and Lipkin, W.I. 2014. Middle East respiratory syndrome coronavirus infection in dromedary camels in Saudi Arabia. mBio. 5(2), 10–128. Alfieri, A.A., Ribeiro, J., de Carvalho Balbo, L., Lorenzetti, E. and Alfieri, A.F. 2018. Dairy calf rearing unit and infectious diseases: diarrhea outbreak by bovine coronavirus as a model for the dispersion of pathogenic microorganisms. Trop. Anim. Health Prod. 50(8), 1937–1940. Ali, M.A., Shehata, M.M., Gomaa, M.R., Kandeil, A., El-Shesheny, R., Kayed, A.S., El-Taweel, A.N., Atea, M., Hassan, N., Bagato, O., Moatasim, Y., Mahmoud, S.H., Kutkat, O., Maatouq, A.M., Osman, A., McKenzie, P.P., Webby, R.J. and Kayali, G. 2017. Systematic, active surveillance for Middle East respiratory syndrome coronavirus in camels in Egypt. Emerg. Micro. Infec. 6(1), 1–7. Al-Mubarak, F., Thwiny, H. and Hizam, M. 2022. Seroprevalence of Orf virus in sheep in Basrah province, Southern Iraq. Bas. J. Vet. 21(S1), 58–66. Al-Mubarak, F.T.M., Najem, H.A. and Thwiny, H.T. 2023. Molecular identification of the common viral respiratory viruses in backyard chickens in Basrah, Southern Iraq. Korean J. Vet. Res. 63(4), e41. Alrodhan, M.A. 2017. Phylogenetic analysis of MERSCoV in human and camels in Iraq. AL-Qadisiyah Med. J. 13(23), 150–159. Al-Shomrani, B., Manee, M., Alharbi, S., Altammami, M., Alshehri, M., Nassar, M., Bakhrebah, M. and Al-Fageeh, M. 2020. Genomic sequencing and analysis of eight camel-derived Middle East Respiratory Syndrome Coronavirus (MERS-CoV) isolates in Saudi Arabia. Viruses 12(6), 611. Ammar, S.S.M., Mokhtaria, K., Tahar, B.B., Amar, A.A., Redha, B.A., Yuva, B., Si Mohamed, H., Abdellatif, N. and Laid, B. 2014. Prevalence of rotavirus (GARV) and coronavirus (BCoV) associated with neonatal diarrhea in calves in western Algeria. Asian Pac. J. Trop. Biomed. 4, S318–S322. Andries, K., Pensaert, M. and Callebaut, P. 2010. Pathogenicity of hemagglutinating encephalomyelitis (vomiting and wasting disease) virus of pigs, using different routes of inoculation. Zentralbl Veterinaermed B. 25(6), 461–468. Boileau, M.J. and Kapil, S. 2010. Bovine coronavirus associated syndromes. Clin. North Am. Food Anim. Pract. 26(1), 123–146. Brandão, P.E., Gregori, F., Richtzenhain, L.J., Rosales, C.A.R., Villarreal, L.Y.B. and Jerez, J.A. 2006. Molecular analysis of Brazilian strains of bovine coronavirus (BCoV) reveals a deletion within the hypervariable region of the S1 subunit of the spike glycoprotein also found in human coronavirus OC43. Arch. Virol. 151(9), 1735–1748. Casanova, L., Rutala, W.A., Weber, D.J. and Sobsey, M.D. 2009. Survival of surrogate coronaviruses in water. Water Res. 43(7), 1893–1898. Chu, D.K.W., Poon, L.L.M., Gomaa, M.M., Shehata, M.M., Perera, R.A.P.M., Abu Zeid, D., El Rifay, A.S., Siu, L.Y., Guan, Y., Webby, R.J., Ali, M.A., Peiris, M. and Kayali, G. 2014. MERS Coronaviruses in dromedary camels, Egypt. Emerg. Infect. Dis. 20(6), 1049–1053. Coleman, C.M. and Frieman, M.B. 2014. Coronaviruses: important emerging human pathogens. Virol. J. 88(10), 5209–5212. Colina, S.E., Serena, M.S., Echeverría, M.G. and Metz, G.E. 2021. Clinical and molecular aspects of veterinary coronaviruses. Virus Res. 1(297), 198382. Corman, V.M., Landt, O., Kaiser, M., Molenkamp, R., Meijer, A., Chu, D.K., Bleicker, T., Brünink, S., Schneider, J., Schmidt, M.L., Mulders, D.G., Haagmans, B.L., van der Veer, B., van den Brink, S., Wijsman, L., Goderski, G., Romette, J.-L., Ellis, J., Zambon, M., Peiris M, Goossens H, Reusken C, Koopmans MP, Drosten, C. 2020. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 25(3), 2000045. Corman, V.M., Müller, M.A., Costabel, U., Timm, J., Binger, T., Meyer, B., Kreher, P., Lattwein, E., Eschbach-Bludau, M., Nitsche, A., Bleicker, T., Landt, O., Schweiger, B., Drexler, J.F., Osterhaus, A.D., Haagmans, B.L., Dittmer, U., Bonin, F., Wolff, T. and Drosten, C. 2012. Assays for laboratory confirmation of novel human coronavirus (hCoV-EMC) infections. Euro Surveill. 17(49), 20334. De Wit, E., Van Doremalen, N., Falzarano, D. and Munster, V.J. 2016. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 14(8), 523–534. Decaro, N., Elia, G., Campolo, M., Desario, C., Mari, V., Radogna, A., Colaianni, M.L., Cirone, F., Tempesta, M. and Buonavoglia, C. 2008. Detection of bovine coronavirus using a TaqMan-based real-time RT-PCR assay. J. Virol. Methods. 151(2), 167–171. Erles, K., Toomey, C., Brooks, H.W. and Brownlie, J. 2003. Detection of a group 2 coronavirus in dogs with canine infectious respiratory disease. Virology 310(2), 216–223. Gossner, C., Danielson, N., Gervelmeyer, A., Berthe, F., Faye, B., Kaasik Aaslav, K., Adlhoch, C., Zeller, H., Penttinen, P. and Coulombier, D. 2016. Human–dromedary camel interactions and the risk of acquiring zoonotic Middle East respiratory syndrome coronavirus infection. Zoonoses Public Health. 63(1), 1–9. Haagmans, B.L., Al Dhahiry, S.H.S., Reusken, C.B.E.M., Raj, V.S., Galiano, M., Myers, R., Godeke, G.-J., Jonges, M., Farag, E., Diab, A., Ghobashy, H., Alhajri, F., Al-Thani, M., Al-Marri, S.A., Al Romaihi, H.E., Al Khal, A., Bermingham, A., Osterhaus, A.D.M.E., AlHajri, M.M. and Koopmans, M.P.G. 2014. Middle East respiratory syndrome coronavirus in dromedary camels: an outbreak investigation. Lancet Infect. Dis.14(2), 140–145. Hasoksuz, M., Hoet, A.E., Loerch, S.C., Wittum, T.E., Nielsen, P.R. and Saif, L.J. 2002. Detection of respiratory and enteric shedding of bovine coronaviruses in cattle in an Ohio feedlot. J. Vet. Diagn. Invest. 14(4), 308–313. Haverkamp, A.-K., Lehmbecker, A., Spitzbarth, I., Widagdo, W., Haagmans, B.L., Segalés, J., Vergara-Alert, J., Bensaid, A., van den Brand, J.M.A., Osterhaus, A.D.M.E. and Baumgärtner, W. 2018. Experimental infection of dromedaries with Middle East respiratory syndrome-coronavirus is accompanied by massive ciliary loss and depletion of the cell surface receptor dipeptidyl peptidase 4. Sci. Rep. 8(1), 9778. Hemida, M.G., Chu, D.K.W., Poon, L.L.M., Perera, R.A.P.M., Alhammadi, M.A., Ng, H., Siu, L.Y., Guan, Y., Alnaeem, A. and Peiris, M. 2014. MERS coronavirus in dromedary camel Herd, Saudi Arabia. Emerg. Infect. Dis. 20(7), 1231. Jeong, J.H., Kim, G.Y., Yoon, S.S., Park, S.J., Kim, Y.J., Sung, C.M., Jang, O.J., Shin, S.S., Koh, H.B., Lee, B.J. and Lee, C.Y. 2005. Detection and isolation of winter dysentery bovine coronavirus circulat in Korea during 2002-2004. J. Vet. Med. Sci. 67(2), 187–189. Johnson, K.K. and Pendell, D.L. 2017. Market impacts of reducing the prevalence of bovine respiratory disease in United States beef cattle feedlots. Front. Vet. Sci. 9(4), 189. Kandeil, A., Gomaa, M., Shehata, M., El-Taweel, A., Kayed, A.E., Abiadh, A., Jrijer, J., Moatasim, Y., Kutkat, O., Bagato, O., Mahmoud, S., Mostafa, A., El-Shesheny, R., Perera, R.A., Ko, R.L., Hassan, N., Elsokary, B., Allal, L., Saad, A., Sobhy H, McKenzie PP, Webby RJ, Peiris M, Ali MA, Kayali, G. 2019. Middle East respiratory syndrome coronavirus infection in non-camelid domestic mammals. Emerg. Micro. Infec. 8(1), 103–108. Kapil, S., Trent, A.M. and Goyal, S.M. 1990. Excretion and persistence of bovine coronavirus in neonatal calves. Arch. Virol. 115, 127–132. Keha, A., Xue, L., Yan, S., Yue, H. and Tang, C. 2019. Prevalence of a novel bovine coronavirus strain with a recombinant hemagglutinin/esterase gene in dairy calves in China. Transbound. Emerg. Dis. 66(5), 1971–1981. Khalafalla, A.I., Lu, X., Al-Mubarak, A.I. A., Dalab, A.H.S., Al-Busadah, K.A.S. and Erdman, D.D. 2015. MERS-CoV in upper respiratory tract and lungs of dromedary camels, Saudi Arabia, 2013–2014. Emerg. Infect. Dis. 21(7), 1153–1158. Kim, E.M., Cho, H.C., Shin, S.U., Park, J. and Choi, K.S. 2022. Prevalence and genetic characterization of bovine coronavirus identified from diarrheic pre-weaned native Korean calves from 2019 to 2021. Infect. Genet. Evol. 1(100), 105263. Kumar, S.S., Rai, R.B., Dhama, K., Ranganath, G.J., Saminathan, M., Wani, M.Y. and Saravanan, R. 2013. Detection of bovine coronavirus in calf diarrheic samples by indirect antigen capture ELISA and RT-PCR. Res. Opin. Anim. Vet. Sci. 3(8), 225–234. Lai, M.M., Liao, C.L., Lin, Y.J. and Zhang, X. 1994. Coronavirus: how a large RNA viral genome is replicated and transcribed. Infec. Age. Dis. 3(2-3), 98–105. Lee, S.H., Kim, H.Y., Choi, E.W. and Kim, D. 2019. Causative agents and epidemiology of diarrhea in Korean native calves. J. Vet. Sci. 20(6), e64. Li, Q., Shah, T., Wang, B., Qu, L., Wang, R., Hou, Y., Baloch, Z. and Xia, X. 2022. Cross-species transmission, evolution and zoonotic potential of coronaviruses. Front. Cell. Infect. Microbiol. 6(12), 1081370. Mansour, K.A., Hasso, S.A. and Al Rodhan, M.N. 2013. Detection of bovine corona virus in some governorate of Iraq. AL-Qadisiya J. Vet. Med. Sci. 12(1), 24–28. Masters, P.S. 2006. The molecular biology of coronaviruses. Adv. Virus. Res. 1(66), 193–292. McIntosh, K., Chao, R.K., Krause, H.E., Wasil, R., Mocega, H.E. and Mufson, M.A. 1974. Coronavirus infection in acute lower respiratory tract disease of infants. J. Infect. Dis. 130(5), 502–507. Mirkena, T., Walelign, E., Tewolde, N., Gari, G., Abebe, G. and Newman, S. 2018. Camel production systems in Ethiopia: a review of literature with notes on MERS-CoV risk factors. Pastoralism 8(1), 30. Mullis, L., Saif, L.J., Zhang, Y., Zhang, X. and Azevedo, M.S. P. 2012. Stability of bovine coronavirus on lettuce surfaces under household refrigeration conditions. Food Microbiol. 30(1), 180–186. Ommeh, S., Zhang, W., Zohaib, A., Chen, J., Zhang, H., Hu, B., Ge, X.-Y., Yang, X.-L., Masika, M., Obanda, V., Luo, Y., Li, S., Waruhiu, C., Li, B., Zhu, Y., Ouma, D., Odendo, V., Wang, L.-F., Anderson, D.E., Lichoti J, Mungube E, Gakuya F, Zhou P, Ngeiywa KJ, Yan B, Agwanda B, Shi, Z.-L. 2018. Genetic evidence of Middle East Respiratory Syndrome Coronavirus (MERS-Cov) and widespread seroprevalence among camels in Kenya. Virol. Sin. 33(6), 484–492. Omrani, A.S., Al-Tawfiq, J.A. and Memish, Z.A. 2015. Middle East respiratory syndrome coronavirus (MERS-CoV): animal to human interaction. Pathog. Glob. Health. 109(8), 354–362. Pal, M., Berhanu, G., Desalegn, C. and Kandi, V. 2020. Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2): an update. Cureus. 12(3), e7423. Park, S.J., Kim, G.Y., Choy, H.E., Hong, Y.J., Saif, L.J., Jeong, J.H., Park, S.I., Kim, H.H., Kim, S.K., Shin, S.S., Kang, M.I. and Cho, K.O. 2007. Dual enteric and respiratory tropisms of winter dysentery bovine coronavirus in calves. Arch. Virol. 152(10), 1885–1900. Raj, V.S., Farag, E.A.B.A., Reusken, C.B.E.M., Lamers, M.M., Pas, S.D., Voermans, J., Smits, S.L., Osterhaus, A.D.M.E., Al-Mawlawi, N., Al-Romaihi, H.E., AlHajri, M.M., El-Sayed, A.M., Mohran, K.A., Ghobashy, H., Alhajri, F., Al-Thani, M., Al-Marri, S.A., El-Maghraby, M.M., Koopmans, M.P.G. and Haagmans, B.L. 2014. Isolation of MERS coronavirus from a dromedary camel, Qatar, 2014. Emerg. Infect. Dis. 20(8), 1339–1342. Reusken, C.B.E.M., Messadi, L., Feyisa, A., Ularamu, H., Godeke, G.-J., Danmarwa, A., Dawo, F., Jemli, M., Melaku, S., Shamaki, D., Woma, Y., Wungak, Y., Gebremedhin, E.Z., Zutt, I., Bosch, B.-J., Haagmans, B.L. and Koopmans, M.P.G. 2014. Geographic distribution of MERS coronavirus among dromedary camels, Africa. Emerg. Infect. Dis. 20(8), 1370–1374. Reusken, C.B.E.M., Schilp, C., Raj, V.S., De Bruin, E., Kohl, R.H.G., Farag, E.A.B.A., Haagmans, B.L., Al-Romaihi, H., Le Grange, F., Bosch, B.-J. and Koopmans, M.P.G. 2016. MERS-CoV infection of alpaca in a region where MERS-CoV is endemic. Emerg. Infect. Dis. 22(6), 1129–1131. Kumar, S. S., Rai, R. B., Dhama, K., Ranganath, G. J., Saminathan, M., Wani, M. Y., and Saravanan, R. 2013. Detection of bovine coronavirus in calf diarrheic samples by indirect antigen capture ELISA and RT-PCR. Res. Opin. Anim. Vet. Sci. 3, 225–234. Saied, A.A., Metwally, A.A., Mohamed, H.M. and Haridy, M.A. 2021. The contribution of bovines to human health against viral infections. ESPR. 28(34), 46999–47023. Saif, L.J. 2010. Bovine respiratory coronavirus. Vet. Clin. N. Am. Food Anim. Pract.26(2), 349–364. Shin, J., Tark, D., Le, V.P., Choe, S., Cha, R.M., Park, G.N., Cho, I.S., Nga, B.T.T., Lan, N.T. and An, D.J. 2019. Genetic characterization of bovine coronavirus in Vietnam. Vir. Gen. 55(3), 415–420. Shirato, K., Nao, N., Matsuyama, S. and Kageyama, T. 2020. Ultra-rapid real-time RT-PCR method for detecting Middle East respiratory syndrome coronavirus using a mobile PCR device, PCR1100. Jpn. J. Infect. Dis. 73(3), 181–186. Siddell, S., Wege, H. and Ter Meulen, V. 1983. The biology of coronaviruses. J. Gen. Virol. 64(4), 761–776. Singasa, K., Songserm, T., Lertwatcharasarakul, P. and Arunvipas, P. 2017. Molecular and phylogenetic characterization of bovine coronavirus virus isolated from dairy cattle in Central Region, Thailand. Trop. Anim. Health Prod. 49, 1523–1529. Su, S., Wong, G., Shi, W., Liu, J., Lai, A.C.K., Zhou, J., Liu, W., Bi, Y. and Gao, G.F. 2016. Epidemiology, genetic recombination and pathogenesis of coronaviruses. Trends Microbiol. 24(6), 490–502. Takiuchi, E., Stipp, D.T., Alfieri, A.F. and Alfieri, A.A. 2006. Improved detection of bovine coronavirus N gene in faeces of calves infected naturally by a semi-nested PCR assay and an internal control. J. Virol. Methods. 131(2), 148–154. Te, N., Ciurkiewicz, M., van den Brand, J.M.A., Rodon, J., Haverkamp, A.-K., Vergara-Alert, J., Bensaid, A., Haagmans, B.L., Baumgartner, W. and Segalés, J. 2022. Middle East respiratory syndrome coronavirus infection in camelids. Vet. Pathol. 59(4), 546–555. Tennant, B.J., Gaskell, R.M. and Gaskell, C.J. 1994. Studies on the survival of canine coronavirus under different environmental conditions. Vet. Microbiol. 42(2-3), 255–259. Thwiny, H.T. 2016. Diagnosis of carrier state of foot and mouth disease virus in vaccinated and unvaccinated cattle by RT-PCR. Bas. J. Vet. 15(2), 30–38. Thwiny, H.T., Al Hamed, T.A. and Nazzal, A.R. 2018. Seroepidemiological study of Middle East respiratory syndrome (MERS) virus infection in Iraqi dromedary camels. Vet. Arh. 88(2), 191–200. Thwiny, H.T., Mansour, F.T. and Al-Taher, H.A. 2015. Genotypic characterization of infectious bronchitis virus from clinically suspected broilers in Basrah, South of Iraq. Int. J. Sci. Tech. 10(4), 54–59. Toftaker, I., Holmøy, I., Nødtvedt, A., Østerås, O. and Stokstad, M. 2017. A cohort study of the effect of winter dysentery on herd-level milk production. J. Dairy. Sci. 100(8), 6483–6493. Van Doremalen, N., Bushmaker, T. and Munster, V.J. 2013. Stability of Middle East respiratory syndrome coronavirus (MERS-CoV) under different environmental conditions. Euro Surveill. 18(38), 20590. van Doremalen, N., Hijazeen, Z.S.K., Holloway, P., Al Omari, B., McDowell, C., Adney, D., Talafha, H.A., Guitian, J., Steel, J., Amarin, N., Tibbo, M., Abu-Basha, E., Al-Majali, A.M., Munster, V.J. and Richt, J.A. 2017. High prevalence of Middle East respiratory coronavirus in young dromedary camels in Jordan. Vector Borne Zoonotic Dis. 17(2), 155–159. Weiss, S.R. and Leibowitz, J.L. 2011. Coronavirus pathogenesis. Adv. Virus. Res. 1(81), 85–164. Weiss, S.R. and Navas-Martin, S. 2005. Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol. Mol. Biol. 69(4), 635–664. Wernery, U., Corman, V.M., Wong, E.Y.M., Tsang, A.K.L., Muth, D., Lau, S.K.P., Khazanehdari, K., Zirkel, F., Ali, M., Nagy, P., Juhasz, J., Wernery, R., Joseph, S., Syriac, G., Elizabeth, S.K., Patteril, N.A.G., Woo, P.C.Y. and Drosten, C. 2015. Acute Middle East respiratory syndrome coronavirus infection in livestock dromedaries, Dubai, 2014. Emerg. Infect. Dis. 21(6), 1019. Yusof, M.F., Eltahir, Y.M., Serhan, W.S., Hashem, F.M., Elsayed, E.A., Marzoug, B.A., Abdelazim, A.S., Bensalah, O.K.A. and Al Muhairi, S.S. 2015. Prevalence of Middle East respiratory syndrome coronavirus (MERS-CoV) in dromedary camels in Abu Dhabi Emirate, United Arab Emirates. Virus Genes 50(3), 509–513. Zhu, Q., Li, B. and Sun, D. 2022. Advances in bovine coronavirus epidemiology. Viruses. 14(5), 1109. Zohaib, A., Saqib, M., Athar, M.A., Chen, J., Sial, A.-R., Khan, S., Taj, Z., Sadia, H., Tahir, U., Tayyab, M.H., Qureshi, M.A., Mansoor, M.K., Naeem, M.A., Hu, B.-J., Khan, B.A., Ujjan, I.D., Li, B., Zhang, W., Luo, Y., Zhu Y, Waruhiu C, Khan I, Yang XL, Sajid MS, Corman VM, Yan B, Shi, Z.-L. 2018. Countrywide survey for MERS-coronavirus antibodies in dromedaries and humans in Pakistan. Virol. Sin. 33(5), 410–417. | ||
| How to Cite this Article |
| Pubmed Style Madhloom IH, Othman RM, Al-bayati HAM. Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Vet. J.. 2025; 15(2): 765-773. doi:10.5455/OVJ.2025.v15.i2.25 Web Style Madhloom IH, Othman RM, Al-bayati HAM. Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. https://www.openveterinaryjournal.com/?mno=226488 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i2.25 AMA (American Medical Association) Style Madhloom IH, Othman RM, Al-bayati HAM. Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Vet. J.. 2025; 15(2): 765-773. doi:10.5455/OVJ.2025.v15.i2.25 Vancouver/ICMJE Style Madhloom IH, Othman RM, Al-bayati HAM. Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Vet. J.. (2025), [cited January 25, 2026]; 15(2): 765-773. doi:10.5455/OVJ.2025.v15.i2.25 Harvard Style Madhloom, I. H., Othman, . R. M. & Al-bayati, . H. A. M. (2025) Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Vet. J., 15 (2), 765-773. doi:10.5455/OVJ.2025.v15.i2.25 Turabian Style Madhloom, Ibrahim Hasan, Rasha Munther Othman, and Hussein Ali Mohammed Al-bayati. 2025. Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Veterinary Journal, 15 (2), 765-773. doi:10.5455/OVJ.2025.v15.i2.25 Chicago Style Madhloom, Ibrahim Hasan, Rasha Munther Othman, and Hussein Ali Mohammed Al-bayati. "Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq." Open Veterinary Journal 15 (2025), 765-773. doi:10.5455/OVJ.2025.v15.i2.25 MLA (The Modern Language Association) Style Madhloom, Ibrahim Hasan, Rasha Munther Othman, and Hussein Ali Mohammed Al-bayati. "Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq." Open Veterinary Journal 15.2 (2025), 765-773. Print. doi:10.5455/OVJ.2025.v15.i2.25 APA (American Psychological Association) Style Madhloom, I. H., Othman, . R. M. & Al-bayati, . H. A. M. (2025) Molecular detection of coronavirus in camelids and bovines using realtime quantitative polymerase chain reaction in Wasit Province, Iraq. Open Veterinary Journal, 15 (2), 765-773. doi:10.5455/OVJ.2025.v15.i2.25 |