| Review Article | ||
Open Vet. J.. 2025; 15(2): 504-518 Open Veterinary Journal, (2025), Vol. 15(2): 504-518 Review Article Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistanceAswin Rafif Khairullah1, Ikechukwu Benjamin Moses2, Sheila Marty Yanestria3, Fidi Nur Aini Eka Puji Dameanti4, Mustofa Helmi Effendi5,6*, John Yew Huat Tang6, Wiwiek Tyasningsih7, Budiastuti Budiastuti8, Muhammad Khaliim Jati Kusala1, Dea Anita Ariani Kurniasih9, Bantari Wisynu Kusuma Wardhani10, Syahputra Wibowo11, Ilma Fauziah Ma’ruf10, Ima Fauziah1, Riza Zainuddin Ahmad1 and Latifah Latifah121Research Center for Veterinary Science, National Research and Innovation Agency (BRIN), Bogor, Indonesia 2Department of Applied Microbiology, Faculty of Science, Ebonyi State University, Abakaliki, Nigeria 3Faculty of Veterinary Medicine, Universitas Wijaya Kusuma Surabaya, Surabaya, Indonesia 4Laboratory of Veterinary Microbiology and Immunology, Faculty of Veterinary Medicine, Universitas Brawijaya, Malang, Indonesia 5Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 6School of Food Industry, Faculty of Bioresources, and Food Industry, Universiti Sultan Zainal Abidin (Besut Campus), Besut, Malaysia 7Division of Veterinary Microbiology, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, Indonesia 8Study Program of Pharmacy Science, Faculty of Health Science, Universitas Muhammadiyah Surabaya, Surabaya, Indonesia 9Research Center for Public Health and Nutrition, National Research and Innovation Agency (BRIN), Bogor, Indonesia 10Research Center for Pharmaceutical Ingredients and Traditional Medicine, National Research and Innovation Agency (BRIN), Bogor, Indonesia. 11Eijkman Research Center for Molecular Biology, National Research and Innovation Agency (BRIN), Bogor, Indonesia. 12Research Center for Animal Husbandry, National Research and Innovation Agency (BRIN), Bogor, Indonesia *Corresponding Author: Mustofa Helmi Effendi. Division of Veterinary Public Health, Faculty of Veterinary Medicine, Universitas Airlangga, Surabaya, East Java, Indonesia. Email: mhelmieffendi [at] gmail.com Submitted: 21/10/2024 Accepted: 02/01/2025 Published: 28/02/2025 © 2025 Open Veterinary Journal
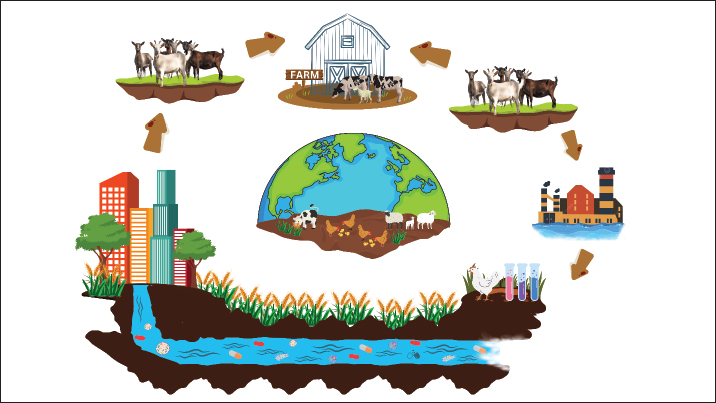
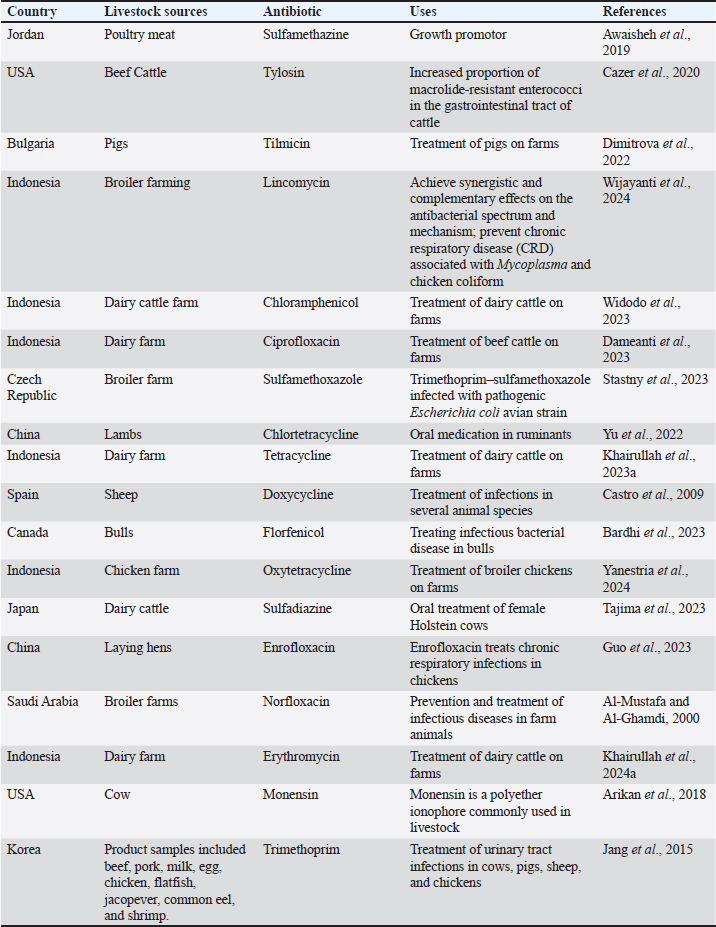
AbstractAntimicrobial resistance (AMR) in bacteria is a global issue requiring serious attention and management. The indiscriminate use of antibiotics in livestock for growth promotion, disease prevention, and treatment has led to the dissemination of AMR bacteria and resistance genes into the environment. In addition, unethical antibiotic sales without prescriptions, poor sanitation, and improper disposal cause significant amounts of antibiotics used in livestock to enter the environment, causing the emergence of resistant bacteria. Intensive livestock farming is an important source of AMR genes, environmental bacteria contamination, and possible transfer to human pathogens. Bacteria intrinsically antibiotic resistant, which are independent of antibiotic use, further complicate AMR and increase the risk of morbidity and mortality following infections by AMR bacteria. Escherichia coli, Salmonella spp., and Staphylococcus spp. are commonly found in livestock that carry resistance genes and have a risk of human infection. The impact of AMR, if left unchecked, could lead to substantial public health burdens globally, with a predicted mortality rate higher than cancer by 2050. “One Health” integrates strategies across human, animal, and environmental health domains, including improving antibiotic stewardship in livestock, preventing infection, and raising awareness regarding the judicious use of antibiotics. The use of antibiotic alternatives, such as prebiotics, probiotics, bacteriophages, bacteriocins, and vaccinations, to control or prevent infections in livestock will help to avoid over-reliance on antibiotics. Coordinated international actions are needed to mitigate the spread of AMR through improved regulations, technology improvements, and awareness campaigns. Keywords: AMR, Antibiotic, Environment, Livestock, Public health. IntroductionEcosystems supporting livestock are crucial sites for the dissemination of antibiotic resistance (Widodo et al., 2022; Ansharieta et al., 2021). The capacity of bacteria to grow and survive in the presence of concentrations of antibacterial drugs that are usually sufficient to either kill or restrict their growth is known as antimicrobial resistance (AMR) (Salam et al., 2023). Additionally, it describes a bacteria’s capacity to fend against the effects of one or more drugs [also known as multidrug resistance (MDR)] (Tyasningsih et al., 2022). Antibiotic pollution in the environment is primarily caused by waste from livestock activities (Kraemer et al., 2019). A significant selection for resistant microbes occurs as a result of up to 90% of antibiotics used in cattle production, which eventually find their way into the environment (Polianciuc et al., 2020). The overuse and improper use of antibiotics over an extended period of time invariably results in the development of bacterial resistance (Khairullah et al., 2020). AMR has become a global concern for people, animals, and the environment in the twenty-first century (Tang et al., 2023). The primary factor contributing to the global spread of AMR is the overuse of antibiotics in the veterinary, medicinal, and agricultural fields (Pradika et al., 2019; Effendi et al., 2018). The problem has been exacerbated by the unethical sale of antibiotics without prescription, inadequate sanitation standards, and the discharge of antibiotic residues or unmetabolized drugs into the environment through industrial waste, manure, and excrement (Serwecińska, 2020). The elements that contribute to the environmental spread of antibiotic resistance are significantly correlated. In the absence of comprehensive action plans, the yearly death toll is predicted to surpass cancer mortality by 2050, with 10 million deaths annually (de Kraker et al., 2016). The degree of antibiotic consumption, which is determined and controlled by a nation’s antibiotic policy, determines the patterns of antibiotic resistance in different areas and nations. Nevertheless, China is recognized as the world’s top manufacturer and user of antibiotics for both humans and animals (Xu et al., 2020). The emergence of AMR in animal products and waste, the abuse of antibiotics that leads to their eventual escape into the environment, and the absence of rigorous and efficient oversight and control over the manufacture, use, and disposal of antibiotics are the main causes of the antibiotic crisis (Muteeb et al., 2023). The emergence and dissemination of antibiotic-resistant genes in food and the environment, together with the abundance of these bacteria, have all been greatly exacerbated by human activities in reaction to industrialization (Manyi-Loh et al., 2018). This also increased the overall abundance of both bacteria and resistance genes. The use of antibiotics in cattle and poultry has recently gained international attention, and it is predicted that by 2050, the percentage of these animals that use antibiotics will reach 67% in developing nations (Rahman et al., 2022). The cost of providing health care worldwide has increased dramatically as a result of the growing AMR issue of a worldwide scale. Although quantifying the global costs of antibiotic resistance is challenging, AMR carries a hefty financial cost (Poudel et al., 2023). The World Health Organization (WHO) has been more interested in funding research to create new antimicrobial compounds against priority pathogens because of the growing resistance of bacteria to antibiotics as a last option (WHO, 2017). Given the lengthy history of antibiotic use in livestock, the establishment of AMR was unavoidable. However, the general public is becoming more aware of sensible measures to reduce unnecessary and overuse of antibiotics. A “One Health” strategy must be used to address AMR because of the interdependence of human, environmental, and animal health (Cella et al., 2023). Understanding how the livestock business environment might serve as a reservoir for the emergence of antibiotic resistance is a crucial topic covered in this article. This review provides fresh perspectives on how to address issues related to AMR in livestock environments. AMR as a global problemAMR, a global problem, is mostly caused by antibiotics and genes that confer resistance to antibiotics. However, other factors such as pollution and poor local sanitation also contribute to AMR (Graham et al., 2019). The main causes of antibiotic resistance are incorrect and insufficient use of antibiotics, ignorance leading to overuse or inappropriate use of antibiotics, poor sanitation and hygiene, excrement releasing unmetabolized antibiotics or their residues into the environment, and the use of antibiotics in poultry and livestock as growth promoters rather than infection control agents (Prestinaci et al., 2015). These factors facilitate the genetic selection pressure that influences the formation of MDR bacterial diseases in the community. The primary pathogens in this category are Salmonella spp., Staphylococcus spp., Escherichia coli, and Campylobacter spp. (Almansour et al., 2023). Furthermore, bacteria from humans or animals have been shown to exhibit similar resistance mechanisms. In the study of resistant microbes, it is crucial to address the molecular evolution of resistance within a given organism, the mechanisms and pathways of transmission between organisms, the spread of resistant microbes between human and animal hosts, and the larger environment, including soil and water (Cycoń et al., 2019). Approximately 80% of the commercially available antibiotics are used as growth promoters or to treat infections in animals (Hosain et al., 2021). The 2010 estimate of livestock antibiotic use on the world antibiotic map was 63,151 tons (Van Boeckel et al., 2015). This predicament has the potential to cost the world economy $120 trillion ($3 trillion annually), or almost the same as the whole yearly budget that the United States currently spends on health care (Aslam et al., 2018). Antibiotic usage patterns will result in 444 million deaths by the year 2100 and a sharp decline in birth rates by the year 2050 (Tang et al., 2023). Use of antibiotics in livestock farmingFor many years, antibiotics have been used in agriculture for growth promotion, prevention, and therapeutic purposes (Van et al., 2020). Presently, the pig, chicken, and cow industries use thousands of tons of antibiotics every year. The most widely used medications include penicillin, chlortetracycline, oxytetracycline, phospholipid flavoring, and bacitracin, according to reports from European countries (EFSA et al., 2021). Many studies have shown that broad-spectrum antibiotics, as listed in Table 1, such as sulfamethazine (Awaisheh et al., 2019), tylosin (Murray et al., 2022), netilmicin (Dimitrova et al., 2022), lincomycin (Wijayanti et al., 2024), chloramphenicol (Widodo et al., 2023), ciprofloxacin (Dameanti et al., 2023), sulfamethoxazole (Stastny et al., 2023), chlortetracycline (Yu et al., 2022), tetracycline (Khairullah et al., 2023a), doxycycline (Castro et al., 2009), florfenicol (Bardhi et al., 2023), oxytetracycline (Yanestria et al., 2024), sulfadiazine (Hahne et al., 2023), enrofloxacin (Guo et al., 2023), norfloxacin (Al-Mustafa and Al-Ghamdi, 2000), erythromycin (Khairullah et al., 2024a), monensin (Arikan et al., 2018), and trimethoprim (Jang et al., 2015) are found in livestock ecosystems across several continents. Genes indicating resistance to tetracycline and macrolide antibiotics are the most commonly identified because of the lengthy history of usage of these antibiotic families in cattle husbandry (Zhuang et al., 2021). Antibiotics are used prophylactically (e.g., to prevent the spread of an infection that is present in a specific area) and therapeutically (e.g., administering antibiotics to the entire herd as a metaphylactic treatment to treat sick animals and prevent the spread of disease) (Rahman et al., 2022). The age and breeding stage of the animal determine the type and quantity of antibiotics used; for example, antibiotics are typically administered during breastfeeding and after weaning. In Belgium, preventive measures account for approximately 90% of the total antibiotic use in pig production (Postma et al., 2016). Antibiotics are currently prohibited from being used as growth promoters in many nations, and attempts are being made to restrict their usage in preventive medicine (Kongsted and Mc Loughlin, 2023). Therefore, the general quantitative consumption levels of these medications were not considerably affected by the prohibition against the use of antibiotics as growth promoters. Antibiotics used in animal agriculture are significantly more common than those used in human medicine. For instance, the Netherlands uses approximately 100 tons of antibiotics annually for the treatment of animals (Mevius and Heederik, 2014). Cross-resistance may develop because many antibiotics used in animal husbandry share structural similarities with those used in human medicine (Wibisono et al., 2022). Livestock as AMR reservoirOne of the reasons for the environmental spread of antibiotic resistance is intensive livestock production (Karwowska, 2024). Livestock facilities are seen as hotspots for antibiotic resistance, and livestock itself is thought to be a significant reservoir of genes and microorganisms resistant to antibiotics (Yanestria et al., 2022). Bacteria with heightened antibiotic resistance can also be found in livestock (Wibisono et al., 2021). Animal isolates may be more likely than human strains to have multiresistant Escherichia coli strains and to be resistant to extended-spectrum β-lactam antibiotics (Xiao et al., 2024). Farm animals are often brought up on cattle and goat farms where antibiotics and their phage are in plentiful supply (Fig. 1). Irrationally excessive antibiotic administration in animal husbandry leads to the acquisition of resistant strains of bacteria (Khairullah et al., 2024b). These resistant bacteria or their genetic material may be disseminated into the environment via several pathways. These pathways include waste materials from farming animals, watersheds, stormwater, and the discharge of antibiotics and resistant bacteria during industrial activity (Kunhikannan et al., 2021). Resistant strains are further distributed to urban and agricultural areas via dirty water (Reddy et al., 2022). This phenomenon perpetuates the cycle of advanced microbial-resistant strains in ecosystems. Urbanization, agricultural development, and industrialization are important factors in the progression of AMR (Mshana et al., 2021). Zhu et al. (2013) found that three large-scale industrial pig farms in China that use antibiotics for farming have 149 distinct genes that cause antibiotic resistance. According to Effendi et al. (2022), the gene blaCTX-M-1 encodes an extended spectrum β-lactamase (ESBL) that is frequently expressed in livestock. It has been discovered that tetracycline resistance genes are present in pig farm animals as well as in the soil or water (Monger et al., 2021). Lau et al. (2017) reported up to 34 novel antibiotic resistance genes in soil samples contaminated with antibiotics used in animal production, such as chlortetracycline, sulfamethazine, and tylosin. The relationship between drug resistance and the use of antibiotics in cattle husbandry is not fully understood. On the one hand, there is no question that the use of antibiotics and the existence of bacteria resistant to the drugs, as well as genes that determine antibiotic resistance, are related (Muteeb et al., 2023). According to studies by Caneschi et al. (2023), using antibiotics in pig production can cause a three- to four-fold increase in the frequency of drug resistance genes. On the other hand, Khairullah et al. (2023b) reported data supporting the presence of antibiotic resistance in pig farms without antibiotic use, suggesting that antibiotic resistance is a naturally occurring source of antibiotic resistance. An unexpected discovery was made by Salerno et al. (2022) that the antibiotic resistance genes blaTEM, sul2, qnrS, and tetA were detected in broiler farms that did not use antibiotics. Their relative abundance was determined to be similar to that of industrial farms as a whole. In addition, studies conducted on chicken farms by Liu et al. (2020a) showed that antibiotic resistance genes were present in broiler farms that used antibiotics as well as those that did not. Table 1. Antibiotic use in livestock farming.
Fig. 1. Pathways of AMR transmission from livestock to the environment.
There is evidence to suggest that certain bacteria in the digestive tracts of animals may naturally withstand antibiotics, independent of the stress brought on by antibiotic use (Kim et al., 2017). Through selection in the presence of antibiotics provided to feed or through contact with drug-resistant microflora in the surrounding environment, such as during grazing, antibiotic-resistant bacteria can colonize livestock species (Xu et al., 2022). This suggests that limiting the use of antibiotics in cattle ranching alone may not be sufficient to curb the spread of drug resistance in livestock. Nonetheless, a correlation has been noted between the antibiotic resistance profile of the bacteria and the makeup and organization of the intestinal microflora in pigs, chickens, and cattle (Wickramasuriya et al., 2022). Antibiotic use as a preventive measure is justified by the fact that intensive farming involves the concentration of a high number of people in a comparatively small area, increasing the risk of infectious disease transmission (Rahman et al., 2022). This is crucial, particularly for large-scale poultry breeding. Concurrently, an Ecuadorian study verified that industrially raised chickens exhibit a higher prevalence of bacterial antibiotic resistance than domestically raised birds (resistance to tetracycline, 78% and 34%, respectively, to sulfisoxazole, 69% and 20%, and to trimethoprim/sulfamethoxazole, 63% and 17%, respectively) (Braykov et al., 2016). Furthermore, it is suggested that the size of the herd, the degree of interpersonal contact, the availability of open areas and pens, and feeding practices all play a role in the development and dissemination of antibiotic resistance in pigs (Redman-White et al., 2023). Österberg et al. (2016) evaluated the prevalence of antibiotic-resistant E. coli bacteria in the digestive tracts of slaughtered pigs from conventional and organic farms in Denmark, Sweden, France, and Italy. The authors discovered that tetracycline, ampicillin, streptomycin, sulfonamides, trimethoprim, ciprofloxacin, nalidixic acid, and gentamicin caused far less antibiotic resistance when farming organically (Iwu et al., 2020). AMR transmission in livestockLivestock excrement contains some of the antibiotics used in livestock production. The body’s ability to metabolize antibiotics varies greatly, from 10% to 90% of the amount taken; this variability is probably influenced by the animal’s age and species (Manyi-Loh et al., 2018). Antibiotics themselves as well as their metabolic byproducts are thus present in the feces. According to a study, animal production solid waste included roughly 12 mg/kg of doxycycline and 241 mg/kg of ciprofloxacin. The antibiotic values for liquid waste were 0.006 and 0.505 mg/l, respectively (Haenni et al., 2022). By comparing these values with concentrations of antibiotics suspected of inducing resistance selection (0.064 µg/l for ciprofloxacin and 2 µg/l for doxycycline), it can be argued that this waste actually threatens the resident microflora’s ability to resist antibiotics (Bengtsson-Palme and Larsson, 2016). Bacteria isolated from animal excrement exhibit antibiotic resistance (Galler et al., 2021). Drug-resistant bacteria are more prevalent in animal feces and the intestinal microflora when antibiotics are administered to feed for therapeutic or prophylactic purposes (Putri et al., 2023). Therefore, it is common to identify agents responsible for antibiotic resistance in animal feces. According to Lima et al. (2020), a number of factors affect the potential role of manure as a hotspot for horizontal gene transfer of traits related to antibiotic resistance, including the abundance of nutrients, the presence of antibiotic residues that may act as selection factors, and the diversity and quantity of microorganisms. Antibiotic resistance genes can be up to 10% more prevalent in manure than 16S rRNA genes. Livestock manure has been found to include a variety of harmful bacteria, including Klebsiella pneumoniae and bacteria with antibiotic resistance traits from the genera Listeria, Salmonella, Coxiella, Campylobacter, and Mycobacterium. The use of manure from livestock farms that have administered antibiotics to their animals as fertilizer is one of the primary drivers of drug resistance in the soil environment (Tian et al., 2021). Research has verified that soil treated with manure contains genes that are resistant to tetracycline, fluoroquinolone, sulfonamide, and chloramphenicol (Wang et al., 2023). In an investigation of antibiotic resistance on Finnish pig and cattle farms, Ruuskanen et al. (2016) discovered that manured soil had high concentrations of resistance genes to tetracyclines (tetM), sulfonamides (sul1), and carbapenems (blaOXA-58), even in situations where animal husbandry typically used very small amounts of antibiotics. It has been noted that germs resistant to antibiotics can endure in the environment for a considerable amount of time. The soil type, composition, frequency of fertilization, and type of microflora are all conditioning factors (Rad et al., 2022). The possibility of additional environmental transmission of antibiotic resistance is another factor. One fascinating phenomenon that may indicate the environmental dissemination of these genes is the discovery of additional antibiotic resistance genes in soil following manure fertilization (Han et al., 2022). Antibiotic resistance in soil microorganisms may be further promoted by antibiotics present in manure (Huygens et al., 2021). The study conducted by Kousar et al. (2021) evaluated the prevalence of antibiotic resistance in Pseudomonas aeruginosa strains isolated from the topsoil of poultry farms where antibiotics were used to grow birds and bacteria of this species from regions at least 500 meters from the closest poultry farm. The samples obtained from both sites showed signs of antibiotic resistance. Fascinatingly, strains of E. coli resistant to florfenicol were identified from farms that were geographically separated. There is evidence that the periodic rise in antibiotic resistance-granting genes in soil environments is caused by manure fertilization (Lima et al., 2020). Additionally, resistance genes might gradually become less common in soil environments. A study by Muurinen et al. (2017) on pig and dairy farms in Finland revealed that although some resistance genes were present in manure, the amount of these genes steadily decreased in soil treated with manure. Baker et al. (2022) found a strikingly similar result for stored manure: a gradual decline in antibiotic-resistant bacteria. It should be noted, nevertheless, that 163 genes related to antibiotic resistance were also found in unfertilized soil, whereas 230–245 genes were found in manured soil, depending on the type of animal. It should be mentioned that using composting is a strategy to lessen the spread of antibiotic resistance caused by the use of natural fertilizers. Antibiotic residues can be removed from samples by up to 50%–99% through manure composting; the greatest results are achieved at higher temperatures and longer thermophilic phases (Li et al., 2020). Regretfully, under composting conditions, certain antibiotics, such as ciprofloxacin, ofloxacin, and sulfamethazine, might not break down while their concentrations in the finished product are still high (Narciso et al., 2023). There is evidence to support the claim that composting manure can effectively reduce the levels of resistance-conditioning genes and bacteria resistant to antibiotics (Huang et al., 2021; Qiu et al., 2021). Composting pig manure at the right temperature and pH can potentially slow the propagation of genes that cause antibiotic resistance, as shown by Liu et al. (2020b). Nevertheless, it was discovered that conventional composting methods were ineffective in managing antibiotic resistance. During the thermophilic phase, even raising the temperature causes a cyclical decrease in the number of resistance-determining genes. Furthermore, our findings suggest that the amount of antibiotic resistance in the composted material may decrease, stabilize, or even grow during composting. Wang et al. (2021) found a nine-fold increase in the number of antibiotic resistance genes (namely, sul1, sul2, tetQ, and tetX genes) in aerobic pile composting of sheep manure. Wang et al. (2022a) reported that the frequency of antibiotic resistance genes increased 44-fold during pile composting, with macrolide antibiotic resistance being the most common type. On the other hand, during composting under thermophilic conditions, tetracycline resistance genes increased to 97%, leading to a 92% decrease in antibiotic resistance genes. Tetracycline resistance gene abundance varies depending on the activity of microflora and environmental conditions (pH, moisture content, and C/N ratio) during composting. The survival of antibiotic-resistant bacteria in manure and compost, as well as the retention of antibiotic resistance genes in microbial communities, may be affected by the presence of heavy metals and drug residues (Shen et al., 2023). Impact of AMR on public healthThe worldwide health care system is financially impacted by the careless discharge of antibiotics into the environment (Polianciuc et al., 2020). The European Center for Disease Control (ECDC, 2018) estimates that 25,000 people die each year from antibiotic-resistant bacterial infections, which affect roughly 2 million Americans annually. The characteristics of AMR include recurrent infections, treatment delays, and resistance spreading to other species (Huemer et al., 2020). Multidrug-resistant health care infections and AMR have given rise to a range of clinical issues. According to the Centers for Disease Control (CDC) estimates, hospitalized patients in the United States have approximately 32,600 cases of MDR Pseudomonas aeruginosa, 197,400 cases of ESBL-producing Enterobacteriaceae, and 223,900 cases of Clostridium difficile annually (CDC, 2019). Given the current situation, more attention should be given to the widespread use of antibiotics and the evaluation of the risks associated with antibiotic residues on human health. Four primary domains, including risk characterization, exposure assessment, hazard identification, and dose–response relationships, should be included in the evaluation of antibiotic-resistant bacterial risks (Murray et al., 2020). Human antibiotic resistance is believed to be caused by changes in the human microbiome caused by antibiotic residues in the environment, which lead to the appearance and selection of resistant bacteria in the human gut (Muteeb et al., 2023). Environmental antibiotic resistance can occasionally result from selective pressure on the environmental microbiome, which serves as a reservoir of antibiotic-resistant bacteria and antimicrobial resistance genes (ARGs) (Larsson and Flach, 2022). In addition to true pathogens such as Haemophilus influenzae, Escherichia coli, Klebsiella pneumoniae, Campylobacter jejuni, and Salmonella spp., Gram-negative pathogens that cause nosocomial infections include Pseudomonas aeruginosa, Clostridium difficile, Burkholderia cepacia, and Acinetobacter baumannii (Davies and Davies, 2010). This infection causes a number of illnesses in humans and animals. According to estimates from the US CDC and Prevention, antibiotic-resistant illnesses claim the lives of 23,000 Americans annually (Thorpe et al., 2018). The first-line antibiotics cannot treat many common diseases, and the germs that cause these infections can also be found in cattle. It is a difficult task to investigate the connections between hazards to human health and the use of antibiotics in livestock husbandry (Bava et al., 2024). There is no straightforward model that explains how resistant microorganisms evolve for epidemiological reasons in terms of the dynamics of antibiotic resistance. It is unknown how much each type of antibiotic contributes to the overall problem of antibiotic resistance. One health approach to AMR managementMicroorganisms are omnipresent and encompass a range of AMR properties in all ecological contexts. AMR between humans, animals, and the environment is becoming more widespread because of the intricate web of interactions that take place between microbial specimens from various “environments,” which promotes gene flow (Palma et al., 2020). This creates a multifaceted issue. Thus, it is preferable to investigate and handle this warning phenomenon using a coordinated multisectoral strategy such as One Health (Velazquez-Meza et al., 2022). To achieve the best possible health for humans, animals, plants, and the environment, One Health is described as “a collaborative effort of multiple health science professions, along with related disciplines and institutions—working locally, nationally, and globally.” (Nzietchueng et al., 2023). According to such a thorough analysis, the primary causes of AMR are the use of antibiotics in the human, animal, and environmental sectors, as well as the global-scale resistance mechanisms that are spreading within and between these sectors (Salam et al., 2023). Because the veterinary and human health care industries share the majority of the same class of drugs, bacteria are subject to cumulative selective pressure, which lowers the effectiveness of antimicrobial-based treatments in the human, veterinary, and environmental domains (Majumder et al., 2020). Increasing awareness when prescribing antibiotic therapy, preventing overprescription, and enhancing hygienic conditions and infection control programs are some of the major steps that the One Health approach has made in the human sector (Muteeb et al., 2023). To lessen the general transmission of AMR features across sectors, One Health initiatives related to the environmental sector include proper handling of industrial, civic, and livestock waste (Chua et al., 2021). Given the complexity and breadth of the One Health approach, the livestock industry should focus on actions fueled by this strategy. In addition to the necessity for international regulations restricting the use of antibiotics as growth promoters in various animal-producing nations, this involves assessing the effects of domestic animal populations and human–animal relationships, the environmental effects of aquaculture, the effectiveness of human and animal treatments, and the reduction of the mass treatment of animal herds (Caneschi et al., 2023). Alternatives to antibioticsThe range of effective substitutes for antibiotics in livestock is largely comparable to that in human medicine. Today, prebiotics and probiotics are widely available; nevertheless, their efficacy is unknown and may vary. It has also been proposed to combine the two, which are referred to as “synbiotics.” The use of bacteriophages and bacteriocins is another alternative (Batista et al., 2020). Bacteriophages exploit bacterial cells for their replication (Secor and Dandekar, 2020). Treatment for Salmonella typhimurium in pigs and poultry has been demonstrated to be beneficial with phage therapy; however, this approach necessitates quick phage delivery and selection (Abd-El Wahab et al., 2023). Two commercial Listeria phage products, Listex P100 and ListeShield, have been approved for use as food preservatives (Kawacka et al., 2020). Numerous studies have investigated how well this new product works against Listeria monocytogenes. For instance, Soni and Nannapaneni (2010) observed a 5-log reduction in L monocytogenes following 24-h, room temperature treatment with Listex P100. Additionally, this substance was tested against developed biofilms of L monocytogenes. Following a 24-h application of Listex P100 at 20°C, Iacumin et al. (2016) observed total disintegration of the biofilm of Listeria monocytogenes on stainless steel wafers. Other authors reported only a 2-log reduction in Listeria during a 2-h treatment period (Sadekuzzaman et al., 2017). One potential antibiotic substitute is the use of bacteriocins, which are antibacterial proteins or peptides generated by ribosomes (Yang et al., 2014). Bacteriocins have attracted considerable interest in the field of antimicrobial research during the past 10 years because of their unique mode of action. These antimicrobial peptides or proteins may have bacteriostatic (i.e., halting cell growth) or bactericidal (i.e., inducing cell death) effects, depending on the type of bacteriocin (Simons et al., 2020). To prevent the synthesis of peptidoglycans, several bacteriocins specifically target lipid II, an integrative molecule found in bacterial cell membranes (Malin and de Leeuw, 2019). Another type of bacteriocin with a binding affinity for lipid II causes the formation of a pore in the bacterial cell membrane. This leads to a reduction in turgor, disruption of electrochemical gradients, and, eventually, cellular demise (Pérez-Ramos et al., 2021). In addition to targeting the bacterial cell wall envelope, several classes of bacteriocins can disrupt essential metabolic processes such as protein synthesis, DNA replication, and gene expression (Darbandi et al., 2020). Lactic acid bacteria (LAB) are the most well-known bacteria that produce bacteriocins, whereas many other species of bacteria can also produce these antimicrobial peptides. Gómez et al. (2016) evaluated the biofilms of three LAB species (Lactococcus lactis, Lactobacillus curvatus, and Lactobacillus sakei) against significant foodborne pathogens, namely L. monocytogenes, E. coli, and Salmonella typhimurium serovar enterica. A different research team by Minei et al. (2008) found a 3.5 log reduction in the attack of L. monocytogenes biofilms within 48 hours after using nisin, a bacteriocin licensed for commercial use, for 9 hours. The variation in bacteriocin is rather high. Alverz-Sieiro et al. (2016) reported that LAB can create over 230 distinct bacteriocins. To some extent, this intriguing antimicrobial peptide has been studied. Nevertheless, none of the aforementioned solutions has widespread commercial availability to treat the entire spectrum of microbial illnesses affecting livestock. Currently, expanding the alternatives for animal immunization might be a more practical strategy. Although vaccines exist for many of the major viral diseases affecting cattle, they are not now routinely used to prevent bacterial infections and diseases (Choudhury et al., 2021). Although the vaccine is not widely used, a trial on live oral Lawsonia inoculation in pigs revealed an 80% reduction in oxytetracycline consumption and enhanced productivity (Bak and Rathkjen, 2009). A long-term plan to reduce antibiotic use in livestock might involve the use of genetically modified animals that are resistant to disease (Mann et al., 2021). An early success in this field is the development of transgenic hens that are unable to spread avian influenza (Lyall et al., 2011). Apart from the previously discussed strategies, there has been positive success in using combinations of molecules to take advantage of the weaknesses of bacteria that would not otherwise respond well to antibiotics. According to Harrison et al. (2019), penicillin can be effective against a considerable proportion of MRSA isolates, including those belonging to the USA300 lineage, when treated with clavulanic acid, a β-lactamase inhibitor. AMR control mitigationNumerous European nations have implemented measures to mitigate the occurrence and dissemination of antibiotic resistance through the prudent administration of antimicrobial medications (Shelke et al., 2023). The US Food and Drug Administration has unveiled a strategy to monitor AMR. AMR has been successfully addressed by nations that have created integrated national programs. The prudent use of antibiotics, the “One Health Approach” for monitoring antibiotics, improvements in health care systems, the creation of health insurance policies, the restriction of medication advertising, well-thought-out disease control guidelines, and community management plans are important variables (Rahman et al., 2022). These tactics also call for a great deal of endurance, time for planning, and adequate government funding. Antibiotic resistance is actively controlled by international organizations such as the Food and Agriculture Organization, the Centers for Disease Control and Prevention, Office International des Epizooties, and the WHO Global AMR Surveillance System. The Global Health Security Agenda (GHSA) and the AMR Action Package (GHSA Prevent-1 Action Package) are two additional initiatives aimed at tackling the worldwide threat of antibiotic resistance. Another pressing issue is diagnostics, particularly in underdeveloped nations where the identification of bacteria is still performed using antiquated microbiology equipment (Wang et al., 2022b). Personalized treatment based on updated and enhanced molecular diagnostic techniques for the necessary antibiotic therapy can close this gap. Research on the interactions between humans and animals and the creation of novel screening instruments can be greatly aided by the One Health approach (Velazquez-Meza et al., 2022). The mechanisms of AMR transmission among all constituents (human, animal, and environment) of a single health system have been uncovered, making this topic extremely relevant and worthy of serious consideration. One of the main causes of AMR, particularly in low- and middle-income countries (LMIC), is the careless and inappropriate use of antibiotics (Otaigbe and Elikwu, 2023). Antibiotics are overprescribed for a number of reasons, including inaccurate diagnoses, especially in poor nations, patient satisfaction with the physician’s prescription, and the bothersome challenges that pharmaceutical businesses face (Thakolkaran et al., 2017). There is an urgent need for advancement in combination therapy, technical innovation, and antibiotic development because the absence of new antibiotics may make it more difficult to investigate this issue (Gupta et al., 2022). AMR and its effects on human and animal health, the role of various factors in its development, the impact of anthropogenic inputs from organizations, and, most importantly, technological, social, and financial initiatives for reducing environmental AMR should be the main areas of focus for future research. ConclusionThere is growing concern about AMR bacteria originating from the livestock industry, which poses a significant threat to humans, animals, and the environment. The livestock industry has become a major reservoir of AMR bacteria and resistance genes due to the indiscriminate use of antibiotics. Resistant pathogens can be transferred to humans through food, water, or direct contact. “One Health” approach highlights the importance of the interconnectedness of human, animal, and environmental health to address AMR. Improved antibiotic stewardship and reduced antibiotic usage in livestock while employing alternatives to promote the growth of good bacteria, such as prebiotics and probiotics, to control pathogen colonization. AMR problems can only be successfully managed through coordinated international action, technology advancement, improved public awareness, and continuous research. Without significant strategies and action to control AMR bacteria, there will be serious consequences in years to come, such as ineffective medical treatment for simple infections. AcknowledgmentsThe authors would like to thank Universitas Airlangga. Conflict of interestThe authors declare no conflict of interest. FundingThis study was partly supported by the International Research Consortium, Lembaga Penelitian dan Pengabdian Masyarakat, Universitas Airlangga, Surabaya, Indonesia, in 2024 (grant number: 171/UN3.LPPM/PT.01.03/2024). Author’s contributionsARK, BWKW, SMY, and SW drafted the manuscript. LL, FNAEPD, MHE, and IBM revised and edited the manuscript. JYHT, DAAK, WT, and IFM participated in preparing and critical checking this manuscript. IF, RZA, BB, and MKJK edited the references. All authors have read and approved the final manuscript. Data availabilityAll references are open-access, so data can be obtained from the online literature. ReferencesAbd-El Wahab, A., Basiouni, S., El-Seedi, H.R., Ahmed, M.F.E., Bielke, L.R., Hargis, B., Tellez-Isaias, G., Eisenreich, W., Lehnherr, H., Kittler, S., Shehata, A.A. and Visscher, C. 2023. An overview of the use of bacteriophages in the poultry industry: successes, challenges, and possibilities for overcoming breakdowns. Front. Microbiol. 14(1), 1136638. Almansour, A.A.M., Alhadlaq, M.M.A., Alzahrani, K.K.O., Mukhtar, L.L.E., Alharbi, A.L. and Alajel, S.M. 2023. Silent threat: antimicrobial-resistant pathogens in food-producing animals and their impact on public health. Microorganisms 11(9), 2127. Al-Mustafa, Z.H. and Al-Ghamdi, M.S. 2000. Use of norfloxacin in poultry production in the eastern province of Saudi Arabia and its potential impact on public health. Int. J. Environ. Health Research. 10(4), 291–299. Alvarez-Sieiro, P., Montalbán-López, M., Mu, D. and Kuipers, O.P. 2016. Bacteriocins of lactic acid bacteria: extending the family. Appl. Microbiol. Biotechnol. 100(7), 2939–2951. Ansharieta, R., Effendi, M.H. and Plumeriastuti, H. 2021. Genetic Identification of Shiga Toxin Encoding Gene from Cases of Multidrug Resistance (MDR) Escherichia coli Isolated from Raw Milk. Trop. Anim. Sci. J. 44(1), 10–15. Arikan, O.A., M’lbry, W., Rice, C. and Lansing, S. 2018. The fate and effect of monensin during anaerobic digestion of dairy manure under mesophilic conditions. PLoS One 13(2), e0192080. Aslam, B., Wang, W., Arshad, M.I., Khurshid, M., Muzammil, S., Rasool, M.H., Nisar, M.A., Alvi, R.F., Aslam, M.A., Qamar, M.U., Salamat, M.K.F. and Baloch, Z. 2018. Antibiotic resistance: a rundown of a global crisis. Infect. Drug Resist. 11(1), 1645–1658. Awaisheh, S.S., Khalifeh, M.S., Rahahleh, R.J., Al-Khaza’leh, J.M. and Algroom, R.M. 2019. sulfamethazine contamination level and exposure assessment in domestic and imported poultry meat in Jordan. World 12(12), 1992–1997. Bak, H. and Rathkjen, P.H. 2009. Reduced use of antimicrobials after vaccination of pigs with porcine proliferative enteropathy in a Danish SPF herd. Acta Vet. Scand. 51(1), 1. Baker, M., Williams, A.D., Hooton, S.P.T., Helliwell, R., King, E., Dodsworth, T., Baena-Nogueras, R.M., Warry, A., Ortori, C.A., Todman, H., Gray-Hammerton, C.J., Pritchard, A.C.W., Iles, E., Cook, R., Emes, R.D., Jones, M.A., Kypraios, T., West, H., Barrett, D.A., Ramsden, S.J., Gomes, R.L., Hudson, C., Millard, A.D., Raman, S., Morris, C., Dodd, C.E.R., Kreft, J.U., Hobman, J.L. and Stekel, D.J. 2022. Antimicrobial resistance in dairy slurry tanks: a critical point for measurement and control. Environ. Int. 169(1), 107516. Bardhi, A., Romano, J.J.E., Pagliuca, G., Caneschi, A. and Barbarossa, A. 2023. Florfenicol and florfenicol amine quantification in bull serum and seminal plasma by a single validated UHPLC-MS/MS method. Vet. Med. Int. 2023(1), 6692920. Batista, V.L., da Silva, T.F., de Jesus, L.C.L., Coelho-Rocha, N.D., Barroso, F.A.L., Tavares, L.M., Azevedo, V., Mancha-Agresti, P. and Drumond, M.M. 2020. Probiotics, prebiotics, synbiotics, and paraprobiotics as therapeutic alternatives for intestinal mucositis. Front. Microbiol. 11(1), 544490. Bava, R., Castagna, F., Lupia, C., Poerio, G., Liguori, G., Lombardi, R., Naturale, M.D., Mercuri, C., Bulotta, R.M., Britti, D. and Palma, E. 2024. Antimicrobial resistance in livestock: a serious threat to public health. Antibiotics 13(6), 551. Bengtsson-Palme, J. and Larsson, D.G.J. 2016. Concentrations of antibiotics predicted to select for resistant bacteria: proposed limits for environmental regulation. Environ. Int. 86(1), 140–149. Braykov, N.P., Eisenberg, J.N.S., Grossman, M., Zhang, L., Vasco, K., Cevallos, W., Muñoz, D., Acevedo, A., Moser, K.A., Marrs, C.F., Foxman, B., Trostle, J., Trueba, G. and Levy, K. 2016. Antibiotic resistance in animal and environmental samples from small-scale poultry farming in northwestern Ecuador. MSphere 10(1), e00021-15. Caneschi, A., Bardhi, A., Barbarossa, A. and Zaghini, A. 2023. The use of antibiotics and antimicrobial resistance in veterinary medicine, a complex phenomenon: a narrative review. Antibiotics 12(3), 487. Castro, L.J., Sahagún, A.M., Diez, M.J., Fernández, N., Sierra, M. and García, J.J., 2009. Pharmacokinetics of doxycycline in sheep after intravenous and oral administration. Vet. J. 180(3), 389–395. CDC. 2019. Antibiotic Resistance Threats In The United States. Atlanta, GA: CDC. Cella, E., Giovanetti, M., Benedetti, F., Scarpa, F., Johnston, C., Borsetti, A., Ceccarelli, G., Azarian, T., Zella, D. and Ciccozzi, M. 2023. Joining forces against antibiotic resistance: the one health solution. Pathogens 12(9), 1074. Choudhury, S.M., Ma, X., Dang, W., Li, Y. and Zheng, H. 2021. Recent development of ruminant vaccine against viral diseases. Front. Vet. Sci. 8(1), 697194. Chua, A.Q., Verma, M., Hsu, L.Y. and Legido-Quigley, H. 2021. An analysis of national action plans for AMR in Southeast Asia using a governance framework approach. Lancet Reg. Health West. Pac. 7(1), 100084. Cycoń, M., Mrozik, A. and Piotrowska-Seget, Z. 2019. Antibiotics in soil environment degradation and their impact on microbial activity and diversity. Front. Microbiol. 10(1), 338. Dameanti, F.N.A.E.P., Yanestria, S.M., Effendi, M.H., Plumeriastuti, H., Tyasningsih, W., Ugbo, E.N., Sutrisno, R. and Safri, M.A.A.S. 2023. Identification of blaSHV and blaTEM extended-spectrum beta-lactamase genes in Klebsiella pneumoniae dairy wastewater from East Java Province, Indonesia. Biodiversitas 24(11), 6092–6099. Darbandi, A., Asadi, A., Ari, M.M., Ohadi, E., Talebi, M., Zadeh, M.H., Emamie, A.D., Ghanavati, R. and Kakanj, M. 2022. Bacteriocins: properties and potential use as antimicrobials. J. Clin. Lab. Anal. 36(1), e24093. Davies, J. and Davies, D. 2010. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74(3), 417–433. De Kraker, M.E., Stewardson, A.J. and Harbarth, S. 2016. Will 10 million people die a year due to antimicrobial resistance by 2050? PLoS Med. 13(11), e1002184. Dimitrova, L., Zaharieva, M. and Najdenski, H. 2022. Pig farms and their surroundings as factors in the spread of antimicrobial resistance. Ecol. Eng. Environ. Prot. 1(2), 14–21. ECDC. 2018. 33000 people die every year because of infections with antibiotic-resistant bacteria. Solna, Sweden: ECDC. Effendi, M.H., Faridah, H.D., Wibisono, F.M., Wibisono, F.J., Nisa, N., Fatimah, F. and Ugbo, E.N. 2022. Detection of genes encoding virulence factors in Escherichia coli isolated from broiler chickens in Blitar District, Indonesia. Biodiversitas 23(7), 3437–3442. Effendi, M.H., Harijani, N., Yanestria, S.M. and Hastutiek, P. 2018. Identification of Shiga toxin-producing Escherichia coli in raw milk samples from dairy cows in Surabaya, Indonesia. Philipp. J. Vet. Med. 55(SI), 109–114. EFSA Panel on Biological Hazards (BIOHAZ), Koutsoumanis, K., Allende, A., Alvarez-Ordóñez, A., Bolton, D., Bover-Cid, S., Chemaly, M., Davies, R., De Cesare, A., Herman, L., Hilbert, F., Lindqvist, R., Nauta, M., Ru, G., Simmons, M., Skandamis, P., Suffredini, E., Andersson, D.I., Bampidis, V., Bengtsson-Palme, J., Bouchard, D., Ferran, A., Kouba, M., Puente, S.L., López-Alonso, M., Nielsen, S.S., Pechová, A., Petkova, M., Girault, S., Broglia, A., Guerra, B., Innocenti, M.L., Liébana, E., López-Gálvez, G., Manini, P., Stella, P. and Peixe, L. 2021. Maximumcross-contamination level forr 24 antimicrobial active substances in nontarget feed. Part 12: tetracyclines: tetracycline, chlortetracycline, oxytetracycline, and doxycycline. EFSA J. 19(10), e06864. Galler, H., Luxner, J., Petternel, C., Reinthaler, F.F., Habib, J., Haas, D., Kittinger, C., Pless, P., Feierl, G. and Zarfel, G. 2021. Multiresistant bacteria isolated from intestinal feces of farm animals in Austria. Antibiotics 10(4), 466. Gómez, N.C., Ramiro, J.M.P., Quecan, B.X.V. and de Melo Franco, B.D.G. 2016. Use of potential probiotic lactic acid bacteria (LAB) biofilms to control Listeria monocytogenes, Salmonella Typhimurium, and Escherichia coli O157: H7 biofilms formation. Front. Microbiol. 7(1), 863. Graham, D.W., Bergeron, G., Bourassa, M.W., Dickson, J., Gomes, F., Howe, A., Kahn, L.H., Morley, P.S., Scott, H.M., Simjee, S., Singer, R.S., Smith, T.C., Storrs, C. and Wittum, T.E. 2019. Complexities in understanding antimicrobial resistance across domesticated animal, human, and environmental systems. Ann. North Y. Acad. Sci. 1441(1), 17–30. Guo, J., Zhang, L., Zhao, Y., Ihsan, A., Wang, X. and Tao, Y. 2023. Metabolic transformation rule of enrofloxacin combined with tilmicosin in laying hens. Metabolites 13(4), 528. Gupta, R. and Sharma, S. 2022. The role of alternatives to antibiotics in mitigating the antimicrobial resistance crisis. Indian J. Med. Res. 156(3), 464–477. Kongsted, H. and McLoughlin, E. T. 2023. Lowering antibiotic uuse and eliminatingpharmaceutical zinc oxide iuse in Danish pig herds: pig farmers’ and veterinarians’ experiences, and perceptions. Livest. Sci. 273(1), 105260. Haenni, M., Dagot, C., Chesneau, O., Bibbal, D., Labanowski, J., Vialette, M., Bouchard, D., Martin-Laurent, F., Calsat, L., Nazaret, S., Petit, F., Pourcher, A.M., Togola, A., Bachelot, M., Topp, E. and Hocquet, D. 2022. Environmental contamination in a high-income country (France) by antibiotics, antibiotic-resistant bacteria, and antibiotic resistance genes: status and possible causes. Environ. Int. 159(1), 107047. Han, B., Ma, L., Yu, Q., Yang, J., Su, W., Hilal, M.G., Li, X., Zhang, S. and Li, H. 2022. Source, fate, and prospect of antibiotic resistance genes in soils: a review. Front. Microbiol. 13(1), 976657. Harrison, E.M., Ba, X., Coll, F., Blane, B., Restif, O., Carvell, H., Köser, C.U., Jamrozy, D., Reuter, S., Lovering, A., Gleadall, N., Bellis, K.L., Uhlemann, A.C., Lowy, F.D., Massey, R.C., Grilo, I.R., Sobral, R., Larsen, J., Larsen, A.R., Lundberg, C.V., Parkhill, J., Paterson, G.K., Holden, M.T.G., Peacock, S.J. and Holmes, M.A. 2019. Genomic identification of cryptic susceptibility to penicillins and β-lactamase inhibitors in methicillin-resistant Staphylococcus aureus. Nat. Microbiol. 4(10), 1680–1691. Hosain, M.M.Z., Kabir, S.M.L. and Kamal, M.M. 2021. Antimicrobial use for livestock production in . Vet. World 14(1), 210–221. Huang, X., Tian, S., Zheng, J., Xu, K. and Liu, C. 2021. Reduced fitness of antibiotic resistome by an extra carbon source during swine manure composting. Environ. Pollut. 277(1), 116819. Huemer, M., Shambat, S.M., Brugger, S.D. and Zinkernagel, A.S. 2020. Antibiotic resistance and persistence—implications for human health and treatment perspectives. EMBO Rep. 21(12), e51034. Huygens, J., Daeseleire, E., Mahillon, J., Van Elst, D., Decrop, J., Meirlaen, J., Dewulf, J., Heyndrickx, M. and Rasschaert, G. 2021. The presence of antibiotic residues and antibiotic-resistant bacteria in cattle manure for fertilization of agricultural fields: a one health perspective. Antibiotics 10(4), 410. Iacumin, L., Manzano, M. and Comi, G. 2016. Phage inactivation of Listeria monocytogenes in San Daniele dry-cured ham and elimination of biofilms from equipment and working environments. Microorganisms 4(1), 4. Iwu, C.D., Korsten, L. and Okoh, A.I. 2020. The incidence of antibiotic resistance within and beyond the agricultural ecosystem: a concern for public health. Microbiol. Open 9(9), e1035. Jang, J.W., Lee, K.S. and Kim, W.S. 2015. Monitoring trimethoprim antibiotic residues in livestock and marine products commercialized in Korea. Food Sci. Biotechnol. 24(6), 1927–1931. Karwowska, E. 2024. Antibiotic resistance in the farming environment. Appl. Sci. 14(13), 5776. Kawacka, I., Olejnik-Schmidt, A., Schmidt, M. and Sip, A. 2020. Effectiveness of phage-based inhibition of Listeria monocytogenes in food products and food processing environments. Microorganisms 8(11), 1764. Khairullah, A.R., Kurniawan, S.C., Effendi, M.H., Sudjarwo, S.A., Ramandinianto, S.C., Widodo, A., Riwu, K.H.P., Silaen, O.S.M. and Rehman, S. 2023b. A review of new livestock-associated methicillin-resistant Staphylococcus aureus from pig farms. World 16(1), 46–58. Khairullah, A.R., Kurniawan, S.C., Sudjarwo, S.A., Effendi, M.H., Afnani, D.A., Silaen, O.S.M., Putra, G.D.S., Riwu, K.H.P., Widodo, A. and Ramandinianto, S.C. 2023a. Detection of multidrug-resistant Staphylococcus aureus and coagulase-negative staphylococci in cow milk from farmers in East Java, Indonesia. Biodiversitas 24(1), 658–664. Khairullah, A.R., Kurniawan, S.C., Sudjarwo, S.A., Effendi, M.H., Widodo, A., Moses, I.B., Hasib, A., Zahra, R.L.A., Gelolodo, M.A., Kurniawati, D.A., Riwu, K.H.P., Silaen, O.S.M., Afnani, D.A. and Ramandinianto, S.C. 2024. Kinship analysis of mecA expression of methicillin-resistant Staphylococcus aureus isolated from milk and risk factors in Blitar, Indonesia. Vet. World 17(1), 216–225. Khairullah, A.R., Sudjarwo, S.A., Effendi, M.H., Kurniawan, S.C., Widodo, A., Silaen, O.S.M. and Ramandinianto, S.C. 2024. Identification of methicillin-resistant Staphylococcus aureus isolated from dairy cow milkin Tulungagung District, Indonesia. Asian J. Dairy Food Res. 43(3), 411–417. Khairullah, A.R., Sudjarwo, S.A., Effendi, M.H., Ramandininto, S.C., Gelolodo, M.A., Widodo, A., Riwu, K.H.P., Kurniawati, D.A. and Rehman, S. 2022. Profile of multidrug resistance and methicillin-resistant Staphylococcus aureus (MRSA) on dairy cows and risk factors from farmer. Biodiversitas 23(6), 2853–2858. Kim, S., Covington, A. and Pamer, E.G. 2017. The intestinal microbiota: antibiotics, colonization resistance, and enteric pathogens. Immunol. Rev. 279(1), 90–105. Kousar, S., Rehman, N., Javed, A., Hussain, A., Naeem, M., Masood, S., Ali, H.A., Manzoor, A., Khan, A.A., Akrem, A., Iqbal, F., Zulfiqar, A., Jamshaid, M.B., Waqas, M., Waseem, A. and Saeed, M.Q. 2021. Intensive poultry farming practices influence antibiotic resistance profiles in Pseudomonas aeruginosa inhabiting nearby soils. Infect. Drug Resist. 14(1), 4511–4516. Kraemer, S.A., Ramachandran, A. and Perron, G.G. 2019. Antibiotic pollution in the environment: from microbial ecology to public policy. Microorganisms 7(6), 180. Kunhikannan, S., Thomas, C.J., Franks, A.E., Mahadevaiah, S., Kumar, S. and Petrovski, S. 2021. Environmental hotspots for antibiotic resistance genes. Microbiologyopen 10(3), e1197. Larsson, D.G.J. and Flach, C.F. 2022. Antibiotic resistance in the environment. Nat. Rev Microbiol. 20(1), 257–269. Lau, C.H., van Engelen, K., Gordon, S., Renaud, J. and Topp, E. 2017. Novel antibiotic resistance determinants in agricultural soil exposed to antibiotics widely used in human medicine and animal farming. Appl. Environ. Microbiol. 83(16), e00989-17. Li, K., Cao, R., Mo, S., Yao, R., Ren, Z. and Wu, J. 2020. Swine manure composting with compound microbial inoculants: removal of antibiotic resistance genes and their associations with microbial community. Front. Microbiol. 11(1), 592592. Lima, T., D’Mingues, S. and Da Silva, G.J. 2020. Manure as a potential hotspot for antibiotic resistance dissemination by horizontal gene transfer events. Vet. Sci. 7(3), 110. Liu, Y., Cheng, D., Xue, J., Weaver, L., Wakelin, S.A., Feng, Y. and Li, Z. 2020b. Changes in microbial community structure during pig manure composting and their relationship to the fate of antibiotics and antibiotic resistance genes. J. Hazard Mater. 389(1), 122082. Liu, Y., Dyall-Smith, M., Marenda, M., Hu, H.W., Browning, G. and Billman-Jacobe, H. 2020a. Antibiotic resistance genes in antibiotic-free chicken farms. Antibiotics 9(3), 120. Lyall, J., Irvine, R.M., Sherman, A., McKinley, T.J., Núñez, A., Purdie, A., Outtrim, L., Brown, I.H., Rolleston-Smith, G. and Sang, H. 2011. Suppression of avian influenza transmission in genetically modified chickens. Science 331(6014), 223–226. Majumder, M.A.A., Rahman, S., Cohall, D., Bharatha, A., Singh, K., Haque, M. and Hilaire, M.G. 2020. Antimicrobial stewardship: fighting antimicrobial resistance and protecting global public health. Infect. Drug Resist. 13(1), 4713–4738. Malin, J.J. and de Leeuw, E. 2019. Therapeutic compounds targeting Lipid II for antibacterial purposes. Infect. Drug Resist. 12(1), 2613–2625. Mann, A., Nehra, K., Rana, J.S. andDahiya, T. 2021. Antibiotic resistance in agriculture: perspectives on upcoming strategies to overcome upsurge in resistance. Curr. Res. Microb. Sci. 2(1), 100030. Manyi-Loh, C., Mamphweli, S., Meyer, E. and Okoh, A. 2018. Antibiotic use in agriculture and its consequential resistance in environmental sources: potential public health implications. Molecules 23(4), 795. Mevius, D. and Heederik, D. 2014. Reduction of antibiotic use in animals “let’s go Dutch”. J. Verbr. Lebensm. 9(1), 177–181. Minei, C.C., Gomes, B.C., Ratti, R.P., D’Angelis, C.E.M. and De Martinis, E.C.P. 2008. Influence of peroxyacetic acid and nisin, coculture with Enterococcus faecium on Listeria monocytogenes biofilm formation. J. Food Prot. 71(3), 634–638. Monger, X.X.C., Gilbert, A.A.A., Saucier, L. and Vincent, A.T. 2021. Antibiotic resistance: from pig to meat. Antibiotics 10(10), 1209. Mshana, S.E., Sindato, C., Matee, M.I. and Mboera, L.E.G. 2021. Antimicrobial use and resistance in african agriculture and food production systems: a systematic review. Antibiotics 10(8), 976. Murray, A.K., Stanton, I.C., Wright, J., Zhang, L., Snape, J. and Gaze, W.H. 2020. The “selection end points in communities of BacTeria” (SELECT) method: a novel experimental assay to facilitate risk assessment of selection for antimicrobial resistance in the environment. Environ. Health Perspect. 128(10), 107007. Murray, S.A., Holbert, A.C., Norman, K.N., Lawhon, S.D., Sawyer, J.E. and Scott, H.M. 2022. Effects of tylosin, a direct-fed microbial and feedlot pen environment, on phenotypic resistance in enterococci isolated from beef cattle feces. Antibiotics 11(1), 106. Muteeb, G., Rehman, M.T., Shahwan, M. and Aatif, M. 2023. Origin of antibiotics and antibiotic resistance and their impacts on drug development: a narrative review. Pharmaceuticals 16(11), 1615. Muurinen, J., Stedtfeld, R., Karkman, A., Parnanen, K., Tiedje, J. and Virta, M. 2017. Influence of manure application on the environmental resistome under Finnish agricultural practice with restricted antibiotic use. Environ. Sci. Technol. 51(11), 5989–5999. Narciso, A., Caracciolo, A.B. and De Carolis, C. 2023. Overview of direct and indirect effects of antibiotics on terrestrial organisms. Antibiotics 12(9), 1471. Nzietchueng, S., Kitua, A., Nyatanyi, T. and Rwego, I.B., 2023. Facilitating implementation of the one-health approach: a definition of a one health intervention. One Health 16(1), 100491. Österberg, J., Wingstrand, A., Jensen, A.N., Kerouanton, A., Cibin, V., Barco, L., Denis, M., Aabo, S. and Bengtsson, B. 2016. Antibiotic resistance in Escherichia coli from pigs in organic and conventional farming in four European Countries. PLoS One 11(6), e0157049. Otaigbe, I.I. and Elikwu, C.J. 2023. Drivers of inappropriate antibiotic use in low- and middle-income countries. JAC Antimicrob. Resist. 5(3), dlad062. Palma, E., Tilocca, B. and Roncada, P. 2020. Antimicrobial resistance in veterinary medicine: an overview. Int. J. Mol. Sci. 21(6), 1914. Pérez-Ramos, A., Madi-Moussa, D., Coucheney, F. and Drider, D., 2021. Current knowledge of the mode of action and immune mechanisms of LAB-bacteriocins. Microorganisms 9(10), 2107. Polianciuc, S.I., Gurzău, A.E., Kiss, B., Ştefan, M.G. and Loghin, F. 2020. Antibiotics in the environment: causes and consequences. Med. Pharm. Rep. 93(3), 231–240. Postma, M., Backhans, A., Collineau, L., Loesken, S., Sjölund, M., Belloc, C., Emanuelson, U., Beilage, E.G., Nielsen, E.O., Stärk, K.D.C., Dewulf, J. and MINAPIG Consortium. 2016. Evaluation of the relationship between biosecurity status, production parameters, herd characteristics, and antimicrobial use in farrow-to-finish pig production in four EU countries. Porcine Health Manag. 2(1), 9. Poudel, A.A.N., Zhu, S., Cooper, N., Little, P., Tarrant, C., Hickman, M. and Yao, G. 2023. The economic burden of antibiotic resistance: a systematic review and meta-analysis. PLoS One 18(5), e0285170. Pradika, A.Y., Chusniati, S., Purnama, M.T.E., Effendi, M.H., Yudhana, A. and Wibawati, P.A. 2019. Total Test of Escherichia coli on Fresh Cow Milk at Dairy Farmer Cooperative (KPSP) Karyo Ngremboko Purwoharjo Banyuwangi. J. Med. Vet. 2(1): 1–6. Prestinaci, F., Pezzotti, P. and Pantosti, A. 2015. Antimicrobial resistance: a global multifaceted phenomenon. Pathog. Glob. Health 109(7), 309–318. Putri, M.F.R., Kendek, I.A., Wibisono, F.J., Effendi, M.H., Rahardjo, D., Tyasningsih, W. and Ugbo, E.N. 2023. Molecular detection of iron in multidrug-resistant avian fecal Escherichia coli isolated from broiler on traditional markets in Surabaya, Indonesia. Biodiversitas 24(12), 6454–6460. Qiu, X., Zhou, G., Chen, L. and Wang, H. 2021. Additive quality influences the reservoir of antibiotic resistance genes during chicken manure composting. Ecotoxicol. Environ. Saf. 220(1), 112413. Rad, A.K., Astaykina, A., Streletskii, R., Afsharyzad, Y., Etesami, H., Zarei, M. and Balasundram, S.K., 2022. An overview of antibiotic resistance and abiotic stresses affecting antimicrobial resistance in agricultural soils. Public Health 19(8), 4666. Rahman, M.M., Tumpa, M.A.A., Zehravi, M., Sarker, M.T., Yamin, M., Islam, M.R., Harun-Or-Rashid, M., Ahmed, M., Ramproshad, S., Mondal, B., Dey, A., Damiri, F., Berrada, M., Rahman, M.H. and Cavalu, S. 2022. An overview of antimicrobial stewardship optimization: the use of antibiotics in humans and animals to prevent resistance. Antibiotics 11(5), 667. Reddy, S., Kaur, K., Barathe, P., Shriram, V., Govarthanan, M. and Kumar, V. 2022. Antimicrobial resistance in urban river ecosystems. Microbiol. Res. 263(1), 127135. Redman-White, C.J., Moran, D., Peters, A.R. and Muwonge, A. 2023. A review of the predictors of antimicrobial use and resistance in European food animal production. Front. Antibiot. 2(1), 1209552. Ruuskanen, M., Muurinen, J., Meierjohan, A., Pärnänen, K., Tamminen, M., Lyra, C., Kronberg, L. and Virta, M. 2016. Fertilizing with animal manure disseminates antibiotic resistance genes to the farm environment. J. Environ. Qual. 45(2), 488–493. Sadekuzzaman, M., Yang, S., Mizan, M.F.R., Kim, H.S. and Ha, S., 2017. Effectiveness of a phage cocktail as a biocontrol agent against L monocytogenes biofilms. Food Control 78(1), 256–263. Salam, M.A., Al-Amin, M.Y., Salam, M.T., Pawar, J.S., Akhter, N., Rabaan, A.A. and Alqumber, M.A.A. 2023. Antimicrobial resistance: a growing serious threat to global public health. Health Care 11(13), 1946. Salerno, B., Furlan, M., Sabatino, R., Di Cesare, A., Leati, M., Volanti, M., Barco, L., Orsini, M., Losasso, C. and Cibin, V. 2022. Antibiotic resistance genes in an antibiotic-free organic broiler farm. Poult. Sci. 101(3), 101675. Secor, P.R. and Dandekar, A.A. 2020. More than simple parasites: the sociobiology of bacteriophages and their bacterial hosts. MBio 11(2), e00041-20. Serwecińska, L. 2020. Antimicrobials and antibiotic-resistant bacteria: a risk to the environment and to public health. Water 12(12), 3313. Shelke, Y.Y.P., Bankar, N.N.J., Bandre, G.G.R., Hawale, D.V. and Dawande, P. 2023. An overview of preventive strategies and the role of various organizations in combating antimicrobial resistance. Cureus 15(9), e44666. Shen, C., He, M., Zhang, J., Liu, J., Su, J. and Dai, J. 2023. Effects of the coexistence of antibiotics and heavy metals on the fate of antibiotic resistance genes in chicken manure and surrounding soils. Ecotoxicol. Environ. Saf. 263(1), 115367. Simons, A., Alhanout, K. and Duval, R.E. 2020. Bacteriocins, antimicrobial peptides of bacterial origin: overview of their biology and their impact against multidrug-resistant bacteria. Microorganisms 8(5), 639. Soni, K.A. and Nannapaneni, R. 2010. Removal of Listeria monocytogenes biofilms with bacteriophage P100. J. Food Prot. 73(8), 1519. Stastny, K., Hodkovicova, N., Jerabek, M., Petren, M., Viskova, M., Papouskova, A., Bartejsova, I., Putecova-Tosnerova, K., Charvatova, M., Zouharova, M., Matiaskova, K. and Nedbalcova, K. 2023. Dosage optimization of trimethoprim and sulfamethoxazole for the treatment of avian pathogenic Escherichia coli in broiler chickens. Antibiotics 13(1), 11. Tajima, T., Saiga, M., Yamamoto, H., Elbadawy, M., Abugomaa, A., Miura, R., Usui, T., Sasaki, K. and Shimoda, M., 2023. Oral pharmacokinetics of sulfadiazine and sulfamonomethoxine in female Holstein milking cows. J. Vet. Med. Sci. 85(7), 715–720. Tang, K.W.K., Millar, B.C. and Moore J.E. 2023. Antimicrobial resistance (AMR). Br. J. Biomed. Sci. 80(1), 11387. Thakolkaran, N., Shetty, A.V., D’Souza, N.D.R. and Shetty, A.K. 2017. Antibiotic prescribing knowledge, attitudes, and practices among physicians in teaching hospitals in South India. J. Family Med. Prim. Care. 6(3), 526–532. Thorpe, K.E., Joski, P. and Johnston, K.J., 2018. Antibiotic-resistant infection treatment costs have doubled since 2002, now exceeding $2 billion annually. Health Aff. 37(4), 662–669. Tian, M., He, X., Feng, Y., Wang, W., Chen, H., Gong, M., Liu, D., Clarke, J.L. and van Eerde, A. 2021. Antibiotic pollution and antimicrobial resistance in livestock and poultry manure in China, and countermeasures. Antibiotics 10(5), 539. Tyasningsih, W., Ramandinianto, S.C., Ansharieta, R., Witaningrum, A.M., Permatasari, D.A., Wardhana, D.K., Effendi, M.H. and Ugbo, E.N. 2022. Prevalence and antibiotic resistance of S. aureus and E. coli isolated from raw milk in East Java, Indonesia. Vet. World 15(8), 2021–2028. Van Boeckel, T.P., Brower, C., Gilbert, M., Grenfell, B.T., Levin, S.A., Robinson, T.P., Teillant, A. and Laxminarayan, R. 2015. Global trends in antimicrobial use in food animals. Proc. Natl. Acad. Sci. U S A 112(18), 5649–5654. Van, T.T.H., Yidana, Z., Smooker, P.M. and Coloe, P.J. 2020. Antibiotic use in food animals worldwide, with a focus on Africa: pluses and minuses. J. Glob. Antimicrob. Resist. 20(1), 170–177. Velazquez-Meza, M.M.E., Galarde-López, M., Carrillo-Quiróz, B. and Alpuche-Aranda, C.M. 2022. Antimicrobial resistance: one health approach. Vet. World 15(3), 743–749. Wang, F., Fu, Y.H., Sheng, H.J., Topp, E., Jiang, X., Zhu, Y.G. and Tiedje, J.M. 2021. Antibiotic resistance in soil ecosystems: a one health perspective. Curr. Opin. Environ. Sci. Health 20(1), 100230. Wang, G., Kong, Y., Yang, Y., Ma, R., Li, L., Li, G. and Yuan, J. 2022a. Composting temperature directly affects the removal of antibiotic resistance genes and mobile genetic elements in livestock manure. Environ. Pollut. 303(1), 119174. Wang, H., Zhang, W. and Tang, Y.W. 2022b. Clinical microbiology in dthe detectionand identification of emerging microbial pathogens: past, present, and future. Emerg. Microbes Infect. 11(1), 2579–2589. Wang, Z., Zhang, N., Li, C. and Shao, L 2023. Diversity of antibiotic resistance genes in soils following four different fertilization treatments. Front. Microbiol. 14(1), 1291599. WHO. 2017. WHO publishes a list of bacteria for which new antibiotics are urgently needed. Geneva: WHO. Wibisono, F.J., Sumiarto, B., Untari, T., Effendi, M.H., Permatasari, D.A. and Witaningrum, A.M. 2021. Molecular identification of the ctx gene of extended-spectrum betalactamase (ESBL) producing Escherichia coli in layer chickens in Blitar, Indonesia. J. Anim. Plant Sci. 31(4), 954–959. Wibisono, F.J., Effendi, M.H. and Wibisono, F.M. 2022. Occurrence, antimicrobial resistance, and potential zoonosis risk of avian pathogenic Escherichia coli in Indonesia: a review. Int. J. One Health 8(2), 76–85. Wickramasuriya, S.S., Park, I., Lee, K., Lee, Y., Kim, W.H., Nam, H. and Lillehoj, H.S. 2022. The role of physiology, immunity, microbiota, and infectious diseases in the gut health of poultry. Vaccines 10(2), 172. Widodo, A., Lamid, M., Effendi, M.H., Khairullah, A.R., Riwu, K.H.P., Yustinasari, L.R., Kurniawan, S.C., Ansori, A.N.M., Silaen, O.S.M. and Dameanti, F.N.A.E.P. 2022. Antibiotic sensitivity profile of multidrug-resistant (MDR) Escherichia coli isolated from dairy cow’s milk in Probolinggo, Indonesia. Biodiversitas 23(10), 4971–4976. Widodo, A., Lamid, M., Effendi, M.H., Tyasningsih, W., Raharjo, D., Khairullah, A.R., Kurniawan, S.C., Yustinasari, L.R., Riwu, K.H.P. and Silaen, O.S.M. 2023. Molecular identification of the blaTEM and blaCTX-M genes in multidrug-resistant Escherichia coli found in milk samples from dairy cattle farms in Tulungagung, Indonesia. J. Vet. Res. 67(3), 381–388. Wijayanti, A.A.D., Muzaki, A.A.Y., Wibisono, C. and Widiasih, D.A. 2024. Therapeutic effects of lincomycin and levels of drug degradation in broiler tissues after treatment. Vet. World 17(5), 1026–1034. Xiao, N., Li, Y., Lin, H., Yang, J., Xiao, G., Jiang, Z., Zhang, Y., Chen, W., Zhou, P., Sun, Z. and Li, J. 2024. Characterization of extended-spectrum β-lactamase-producing Escherichia coli in animal farms in Hunan Province, China. Microorganisms 12(4), 653. Xu, C., Kong, L., Liao, Y., Tian, Y., Wu, Q., Liu, H. and Wang, X. 2022. Mini-review: antibiotic-resistant Escherichia coli from farm animal-associated sources. Antibiotics 11(11), 1535. Xu, J., Sangthong, R., McNeil, E., Tang, R. and Chongsuvivatwong, V. 2020. Antibiotic use in chicken farms in northwestern China. Infect. Control 9(1), 10. Yanestria, S.M., Dameanti, F.N.A.E.P., Musayannah, B.G., Pratama, J.W.A., Witaningrum, A.M., Effendi, M.H., and Ugbo, E.N. 2022. Antibiotic resistance pattern of Extended-Spectrum β-Lactamase (ESBL) producing Escherichia coli isolated from a broiler farm environment in Pasuruan district, Indonesia. Biodiversitas 23(9), 4460–4465. Yanestria, S.M., Effendi, M.H., Tyasningsih, W., Moses, I.B., Khairullah, A.R., Kurniawan, S.C., Dameanti, F.N.A.E.P., Ikaratri, R., Pratama, J.W.A., Sigit, M., Hasib, A. and Silaen, O.S.M. 2024. Antimicrobial resistance patterns and genes of Campylobacter jejuni isolated from chickens in Pasuruan, Indonesia. Open Vet. J. 14(3), 759–768. Yang, S.C., Lin, C.H., Sung, C.T. and Fang, J.Y. (2014). Antibacterial activities of bacteriocins: application in foods and pharmaceuticals. Front. Microbiol. 5(1), 241. Yu, Y., Li, X., Liu, Z., Xu, Y., Shen, Y., Li, G. and Huang, X. 2022. Effects of chlortetracycline-protected granules on rumen microorganisms and their diarrhea therapeutic effect. Front. Vet. Sci. 9(1), 840442. Zhu, Y.G., Johnson, T.A., Su, J.Q., Qiao, M., Guo, G.X., Stedtfeld, R.D., Hashsham, S.A. and Tiedje, J.M. 2013. Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc. Natl. Acad. Sci. U S A 110(9), 3435–3440. Zhuang, M., Achmon, Y., Cao, Y., Liang, X., Chen, L., Wang, H., Siame, B.A. and Leung, K.Y. 2021. Distribution of antibiotic resistance genes in the environment. Environ. Pollut. 285(1), 117402. | ||
| How to Cite this Article |
| Pubmed Style Khairullah AR, Moses IB, Yanestria SM, , Effendi MH, Tang JYH, Tyasningsih W, Budiastuti B, Kusala MKJ, Kurniasih DAA, Wardhani BWK, Wibowo S, Ma'ruf IF, Fauziah I, Ahmad RZ, Latifah L. Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Vet. J.. 2025; 15(2): 504-518. doi:10.5455/OVJ.2025.v15.i2.2 Web Style Khairullah AR, Moses IB, Yanestria SM, , Effendi MH, Tang JYH, Tyasningsih W, Budiastuti B, Kusala MKJ, Kurniasih DAA, Wardhani BWK, Wibowo S, Ma'ruf IF, Fauziah I, Ahmad RZ, Latifah L. Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. https://www.openveterinaryjournal.com/?mno=225539 [Access: January 25, 2026]. doi:10.5455/OVJ.2025.v15.i2.2 AMA (American Medical Association) Style Khairullah AR, Moses IB, Yanestria SM, , Effendi MH, Tang JYH, Tyasningsih W, Budiastuti B, Kusala MKJ, Kurniasih DAA, Wardhani BWK, Wibowo S, Ma'ruf IF, Fauziah I, Ahmad RZ, Latifah L. Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Vet. J.. 2025; 15(2): 504-518. doi:10.5455/OVJ.2025.v15.i2.2 Vancouver/ICMJE Style Khairullah AR, Moses IB, Yanestria SM, , Effendi MH, Tang JYH, Tyasningsih W, Budiastuti B, Kusala MKJ, Kurniasih DAA, Wardhani BWK, Wibowo S, Ma'ruf IF, Fauziah I, Ahmad RZ, Latifah L. Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Vet. J.. (2025), [cited January 25, 2026]; 15(2): 504-518. doi:10.5455/OVJ.2025.v15.i2.2 Harvard Style Khairullah, A. R., Moses, . I. B., Yanestria, . S. M., , Effendi, . M. H., Tang, . J. Y. H., Tyasningsih, . W., Budiastuti, . B., Kusala, . M. K. J., Kurniasih, . D. A. A., Wardhani, . B. W. K., Wibowo, . S., Ma'ruf, . I. F., Fauziah, . I., Ahmad, . R. Z. & Latifah, . L. (2025) Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Vet. J., 15 (2), 504-518. doi:10.5455/OVJ.2025.v15.i2.2 Turabian Style Khairullah, Aswin Rafif, Ikechukwu Benjamin Moses, Sheila Marty Yanestria, Fidi Nur Aini Eka Puji Dameanti, Mustofa Helmi Effendi, John Yew Huat Tang, Wiwiek Tyasningsih, Budiastuti Budiastuti, Muhammad Khaliim Jati Kusala, Dea Anita Ariani Kurniasih, Bantari Wisynu Kusuma Wardhani, Syahputra Wibowo, Ilma Fauziah Ma'ruf, Ima Fauziah, Riza Zainuddin Ahmad, and Latifah Latifah. 2025. Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Veterinary Journal, 15 (2), 504-518. doi:10.5455/OVJ.2025.v15.i2.2 Chicago Style Khairullah, Aswin Rafif, Ikechukwu Benjamin Moses, Sheila Marty Yanestria, Fidi Nur Aini Eka Puji Dameanti, Mustofa Helmi Effendi, John Yew Huat Tang, Wiwiek Tyasningsih, Budiastuti Budiastuti, Muhammad Khaliim Jati Kusala, Dea Anita Ariani Kurniasih, Bantari Wisynu Kusuma Wardhani, Syahputra Wibowo, Ilma Fauziah Ma'ruf, Ima Fauziah, Riza Zainuddin Ahmad, and Latifah Latifah. "Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance." Open Veterinary Journal 15 (2025), 504-518. doi:10.5455/OVJ.2025.v15.i2.2 MLA (The Modern Language Association) Style Khairullah, Aswin Rafif, Ikechukwu Benjamin Moses, Sheila Marty Yanestria, Fidi Nur Aini Eka Puji Dameanti, Mustofa Helmi Effendi, John Yew Huat Tang, Wiwiek Tyasningsih, Budiastuti Budiastuti, Muhammad Khaliim Jati Kusala, Dea Anita Ariani Kurniasih, Bantari Wisynu Kusuma Wardhani, Syahputra Wibowo, Ilma Fauziah Ma'ruf, Ima Fauziah, Riza Zainuddin Ahmad, and Latifah Latifah. "Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance." Open Veterinary Journal 15.2 (2025), 504-518. Print. doi:10.5455/OVJ.2025.v15.i2.2 APA (American Psychological Association) Style Khairullah, A. R., Moses, . I. B., Yanestria, . S. M., , Effendi, . M. H., Tang, . J. Y. H., Tyasningsih, . W., Budiastuti, . B., Kusala, . M. K. J., Kurniasih, . D. A. A., Wardhani, . B. W. K., Wibowo, . S., Ma'ruf, . I. F., Fauziah, . I., Ahmad, . R. Z. & Latifah, . L. (2025) Potential of the livestock industry environment as a reservoir for spreading antimicrobial resistance. Open Veterinary Journal, 15 (2), 504-518. doi:10.5455/OVJ.2025.v15.i2.2 |